Bulletin of Botanical Research ›› 2026, Vol. 46 ›› Issue (2): 231-245.doi: 10.7525/j.issn.1673-5102.2026.02.004
• Original Paper • Previous Articles Next Articles
Yijiao LIU1,2, Jiqing ZHAO1,2, Jinhuan CHEN1,2( )
)
Received:2026-01-07
Online:2026-03-20
Published:2026-04-02
Contact:
Jinhuan CHEN
E-mail:chenjh@bjfu.edu.cn
CLC Number:
Yijiao LIU, Jiqing ZHAO, Jinhuan CHEN. Cloning and Functional Study of the LrMYB64 Gene in Lyciumruthenicum[J]. Bulletin of Botanical Research, 2026, 46(2): 231-245.
Add to citation manager EndNote|Ris|BibTeX
URL: https://bbr.nefu.edu.cn/EN/10.7525/j.issn.1673-5102.2026.02.004
Table 1
Application of the primer sequences
引物名称 Primer name | 引物序列(5′→3′) Primer sequence(5′→3′) | 用途 Application |
|---|---|---|
| LrMYB64-F | ATGAGAAAACCTTGTTGTGATCA | 全长克隆引物序列 |
| LrMYB64-R | TTAAGGAAGTGAGTTGAGATCAGGC | 全长克隆引物序列 |
| 35s-F | GACGCACAATCCCACTATCC | 检测 |
| 35s-R | AATCATCGCAAGACCGGC | 检测 |
| LrMYB64-接头-F | GCGGGTCGACGGTACCATGAGAAAACCTTGTTGTG | 重组引物 |
| LrMYB64-接头-R | TAGACATATGGGTACCTTAAGGAAGTGAGTTGAGAT | 重组引物 |
Table 4
RT-qPCR primers sequence
引物名称 Primer name | 引物序列(5′→3′) Primer sequence(5′→3′) | 引物名称 Primer name | 引物序列(5′→3′) Primer sequence(5′→3′) |
|---|---|---|---|
| Actin-7-F | CTGACGGTGAGGACATTCA | LrDFR-qPCR-F | AGGCTGCAATGGAAGAAGCT |
| Actin-7-R | GAGCATCATCTCCAGCAAAG | LrDFR-qPCR-R | AGCTAGGTGGGAACGTAGGT |
| LrFLS1-qPCR-F | TTGTCACTTGGGCTTGGGTT | LrCHI3-qPCR-F | CAGCAGTGGAAGGGCAAAAC |
| LrFLS1-qPCR-R | AAATCAGGCCTGGGACATGG | LrCHI3-qPCR-R | TGCACCCCATACTGTGAACC |
| LrF3H-qPCR-F | AAGGCATGTGAGGATTGGGG | LrPP2C51-qPCR-F | AGAAAGTCTCGCCACACAGG |
| LrF3H-qPCR-R | CAGTCTTGGACCACTTCCCC | LrPP2C51-qPCR-R | TTCCTCAACCAAGGTGTCGG |
| LrMYB64-qPCR-F | ACATGGTGAAGATTGCTGGAGA | LrGH3.6-like2-qPCR-F | TGTGCTTCGAGTTGCTGGAT |
| LrMYB64-qPCR-R | CTTCCAGCTATTAGGGACCATCT | LrGH3.6-like2-qPCR-R | GCACTGCATTGTTCACTGCA |
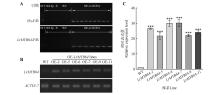
Fig.4
Identification of positive plants with OE-LrMYB64 gene of Lycium ruthenicumA. Identificationat the DNA level; B. RT-PCR gel electrophoresis image; C. The relative expression levels of the OE-LrMYB64 lines. ***.Significant difference in the relative gene expression between transgenic plants and wild type at the 0.001 level. WT. Wild lines; OE. Gene-overexpressed lines.
Table 6
Summary of RNA-seq data output and quality for WT and OE-LrMYB64 lines of Lyciumruthenicum
样品 Sample | 过滤后读段数 Clean reads/M | 过滤后碱基数 Clean bases/Gb | GC比例 GC percentage/% | Q30碱基比例 Q30 base percentage/% |
|---|---|---|---|---|
| WT1 | 22.66 | 6.76 | 42.72 | 98.52 |
| WT2 | 26.35 | 7.84 | 42.86 | 98.53 |
| WT3 | 21.44 | 6.38 | 42.72 | 98.68 |
| OE-LrMYB64-2 | 20.84 | 6.22 | 42.34 | 98.39 |
| OE-LrMYB64-6 | 21.21 | 6.33 | 42.47 | 98.47 |
| OE-LrMYB64-7 | 21.12 | 6.30 | 42.50 | 98.54 |
Table 7
Statistics of RNA-seq reads alignment to the Lyciumruthenicum reference genome
样品 Sample | 总读段数 Total reads/M | 比对读段数 Mapped reads/M | 比对率 Mapped ratio/% | 唯一比对读段数 Uniquely mapped reads/M | 唯一比对率 Uniquely mapped ratio/% |
|---|---|---|---|---|---|
| WT1 | 45.32 | 43.26 | 95.46 | 41.91 | 92.47 |
| WT2 | 52.70 | 50.21 | 95.29 | 48.37 | 91.79 |
| WT3 | 42.87 | 40.76 | 95.06 | 39.33 | 91.72 |
| OE-LrMYB64-2 | 41.68 | 39.07 | 93.74 | 37.84 | 90.78 |
| OE-LrMYB64-6 | 42.42 | 39.95 | 94.17 | 38.63 | 91.06 |
| OE-LrMYB64-7 | 42.25 | 40.16 | 95.05 | 38.85 | 91.96 |
Table 8
Differential expression analysis of key genes in the anthocyanin and flavonoid biosynthesis pathways in OE-LrMYB64 lines
基因名称 Gene name | 基因编号 Gene ID | 基因注释 (NR) Gene annotation (NR) | log2(倍数变化) log2(FC) | 校正后P值 FDR | 上下调情况 Regulation |
|---|---|---|---|---|---|
| LrCHS1B | Lr09g000692 | 查尔酮合酶 Chalcone synthase, partial | -4.720 | 1.44×109 | 下调 |
| LrFLS1 | Lr02g004007 | 黄酮醇合酶/黄烷酮3-羟化酶 Flavonol synthase/flavanone 3-hydroxylase | -4.517 | 3.04×1011 | 下调 |
| LrCHS2 | Lr05g000433 | 查尔酮合酶 Chalcone synthase, partial | -4.312 | 5.40×109 | 下调 |
| LrF3'5'H1 | Lr06g000237 | 黄烷酮-3',5'-羟化酶 Flavanone-3'5'-hydroxylase | -3.438 | 2.05×103 | 下调 |
| LrF3H | Lr12g002886 | 黄烷酮-3-羟化酶 Flavanone-3-hydroxylase | -3.429 | 8.58×106 | 下调 |
| LrA3G2XYLT | Lr01g000190 | UDP-糖基转移酶 79B6 UDP-glycosyltransferase 79B6 | -3.739 | 8.96×103 | 下调 |
| LrDFR | Lr12g002988 | 二氢黄酮醇-4-还原酶 Dihydroflavonol-4-reductase | -3.022 | 7.13×105 | 下调 |
| LrANS | Lr04g003932 | 花青素合酶 Anthocyanidin synthase | -2.882 | 8.86×103 | 下调 |
| LrCHI3 | Lr05g000667 | 查尔酮异构酶 Chalcone isomerase | -2.19 | 4.18×1015 | 下调 |
| LrA3G2XYLT-like | Lr10g003569 | 花青素3-O-葡萄糖苷2‴-O-木糖基转移酶样蛋白 Anthocyanidin 3-O-glucoside 2‴-O-xylosyltransferase-like | -2.053 | 9.54×105 | 下调 |
| LrA3GT | Lr08g002945 | 花青素3-O-葡萄糖基转移酶样前体 Anthocyanidin 3-O-glucosyltransferase-like precursor | -1.940 | 1.44×1013 | 下调 |
| LrPAL-like | Lr03g000391 | 苯丙氨酸解氨酶 Phenylalanine ammonia-lyase | -1.889 | 2.52×108 | 下调 |
| LrPAL | Lr03g000392 | 苯丙氨酸解氨酶 Phenylalanine ammonia-lyase | -1.626 | 1.91×1019 | 下调 |
Table 9
Key differentially expressed genes in the plant hormone signaling pathway of OE-LrMYB64 lines
基因名称 Gene name | 基因编号 Gene ID | 基因注释 (NR) Gene annotation (NR) | log2(倍数变化) log2(FC) | 校正后P值 FDR | 上下调情况 Regulation |
|---|---|---|---|---|---|
| LrPR4 | Lr06g004329 | 病程相关叶片蛋白4样蛋白 Pathogenesis-related leaf protein 4-like | 4.450 | 2.81×104 | 上调 |
| LrPP2C51 | Lr03g002599 | 蛋白磷酸酶2C 51 Probable protein phosphatase 2C 51 | 2.627 | 5.85×103 | 上调 |
| LrSAUR36 | Lr06g003114 | 生长素响应蛋白SAUR36样蛋白 Auxin-responsive protein SAUR36-like | 2.325 | 4.14×105 | 上调 |
| LrGH3.6-like2 | Lr01g000120 | 吲哚-3-乙酸-酰胺合成酶GH3.6样蛋白 Indole-3-acetic acid-amido synthetase GH3.6-like | 2.130 | 5.91×1020 | 上调 |
| LrARF9 | Lr09g004091 | 生长素响应因子9样蛋白 Auxin response factor 9-like | 2.080 | 4.36×103 | 上调 |
| LrGH3.6-like1 | Lr01g000121 | 吲哚-3-乙酸-酰胺合成酶GH3.6样蛋白 Indole-3-acetic acid-amido synthetase GH3.6-like | 1.997 | 5.25×108 | 上调 |
| LrGID1B | Lr01g004492 | 赤霉素受体GID1B样蛋白 Gibberellin receptor GID1B-like | 1.733 | 3.00×103 | 上调 |
| LrSAUR50 | Lr09g003793 | 生长素响应蛋白SAUR50样蛋白 Auxin-responsive protein SAUR50-like | 1.526 | 3.26×103 | 上调 |
| LrABF4 | Lr08g001908 | 脱落酸不敏感5样蛋白4 ABSCISIC ACID-INSENSITIVE 5-like protein 4 | 1.287 | 1.07×108 | 上调 |
| LrGH3.6 | Lr01g000122 | 吲哚-3-乙酸-酰胺合成酶GH3.6样蛋白 Indole-3-acetic acid-amido synthetase GH3.6-like | 1.245 | 1.63×1011 | 上调 |
| LrIAA26 | Lr03g004407 | 生长素响应蛋白IAA26 Auxin-responsive protein IAA26 | 1.155 | 1.50×106 | 上调 |
| LrARF5 | Lr02g005512 | 生长素响应因子5 Auxin response factor 5 | 1.096 | 8.88×109 | 上调 |
| [1] | 杨慧勤,王佳丽,李思蕤,等.茄科蔬菜花青素苷分子调控研究进展[J].生物工程学报,2022,38(5):1738-1752. |
| YANG H Q, WANG J L, LI S R,et al.Advances in the molecular regulation of anthocyanins in solanaceous vegetables[J].Chinese Journal of Biotechnology,2022,38(5):1738-1752. | |
| [2] | 于洋,朱月,刘晗,等.茄果类蔬菜花青素研究进展[J].贵州农业科学,2021,49(8):120-127. |
| YU Y, ZHU Y, LIU H,et al.Research progress on anthocyanins in solanaceous vegetables[J].Guizhou Agricultural Sciences,2021,49(8):120-127. | |
| [3] | JEZ J M, NOEL J P.Mechanism of chalcone synthase.pKa of the catalytic cysteine and the role of the conserved histidine in a plant polyketide synthase[J].Journal of Biological Chemistry,2000,275(50):39640-39646. |
| [4] | 吴榕,田再民,郑国华,等.白三叶CHS基因的过表达提高烟草类黄酮的含量[J].草业科学,2020,37(2):305-313. |
| WU R, TIAN Z M, ZHENG G H,et al.Overexpression of CHS genes from Trifolium repens increases tobacco flavonoid content[J].Pratacultural Science,2020,37(2):305-313. | |
| [5] | 宋天晓,苏文瑾,刘意,等.甘薯黄烷酮3-羟化酶基因IbF3H的克隆和表达特性分析[J].热带作物学报,2021,42(6):1531-1538. |
| SONG T X, SU W J, LIU Y,et al.Cloning and expression analysis of sweetpotato flavanone 3-hydroxylase gene IbF3H [J].Chinese Journal of Tropical Crops,2021,42(6):1531-1538. | |
| [6] | 吴万亿,张毅,张洁,等.马铃薯LINC4206及其靶基因StDFR的克隆与StDFR的生物信息学分析[J].山西农业科学,2021,49(6):683-688. |
| WU W Y, ZHANG Y, ZHANG J,et al.Cloning of LINC4206 and its target gene StDFR and bioinformatics analysis of StDFR in potato[J].Journal of Shanxi Agricultural Sciences,2021,49(6):683-688. | |
| [7] | 姚建忠.毛白杨花青素合酶基因的克隆与特性分析[J].中国细胞生物学学报,2018,40(3):349-356. |
| YAO J Z.Molecular cloning and characterization of two genes encoding anthocyanin synthase from Populus tomentosa [J].Chinese Journal of Cell Biology,2018,40(3):349-356. | |
| [8] | 常博,马长青,刘聪,等. MdMYB1启动子甲基化对不同色泽类型苹果品种果皮花青苷合成的调控作用[J].分子植物育种,2018,16(22):7415-7422. |
| CHANG B, MA C Q, LIU C,et al.Regulation of MdMYB1 promoter methylation on anthocyanin synthesis in peel of apple (Malus×domestica Borkh.) cultivars with different colors[J].Molecular Plant Breeding,2018,16(22):7415-7422. | |
| [9] | AN X H, TIAN Y, CHEN K Q,et al. MdMYB9 and MdMYB11 are involved in the regulation of the JA-induced biosynthesis of anthocyanin and proanthocyanidin in apples[J].Plant and Cell Physiology,2015,56(4):650-662. |
| [10] | LI P, LI Y J, ZHANG F J,et al.The Arabidopsis UDP-glycosyltransferases UGT79B2 and UGT79B3,contribute to cold,salt and drought stress tolerance via modulating anthocyanin accumulation[J].The Plant Journal,2017,89(1):85-103. |
| [11] | 王华,李茂福,杨媛,等.果实花青素生物合成分子机制研究进展[J].植物生理学报,2015,51(1):29-43. |
| WANG H, LI M F, YANG Y,et al.Recent advances on the molecular mechanisms of anthocyanin synthesis in fruits[J].Plant Physiology Journal,2015,51(1):29-43. | |
| [12] | HOLTON T A, CORNISH E C.Genetics and biochemistry of anthocyanin biosynthesis[J].The Plant Cell,1995,7(7):1071-1083. |
| [13] | GONZALI S, MAZZUCATO A, PERATA P.Purple as a tomato:towards high anthocyanin tomatoes[J].Trends in Plant Science,2009,14(5):237-241. |
| [14] | AN X H, TIAN Y, CHEN K Q,et al.The apple WD40 protein MdTTG1 interacts with bHLH but not MYB proteins to regulate anthocyanin accumulation[J].Journal of Plant Physiology,2012,169(7):710-717. |
| [15] | QUATTROCCHIO F, VERWEIJ W, KROON A,et al.PH4 of Petunia is an R2R3 MYB protein that activates vacuolar acidification through interactions with basic-helix-loop-helix transcription factors of the anthocyanin pathway[J].The Plant Cell,2006,18(5):1274-1291. |
| [16] | KOES R, VERWEIJ W, QUATTROCCHIO F.Flavonoids:a colorful model for the regulation and evolution of biochemical pathways[J].Trends in Plant Science,2005,10(5):236-242. |
| [17] | 柯玉洁,陈明堃,马山虎,等.兰科植物MYB转录因子研究进展[J].园艺学报,2021,48(11):2311-2320. |
| KE Y J, CHEN M K, MA S H,et al.Research progress of MYB transcription factors in Orchidaceae[J].Acta Horticulturae Sinica,2021,48(11):2311-2320. | |
| [18] | MATÍAS-HERNÁNDEZ L, JIANG W M, YANG K,et al.AaMYB1 and its orthologue AtMYB61 affect terpene metabolism and trichome development in Artemisia annua and Arabidopsis thaliana [J].The Plant Journal,2017,90(3):520-534. |
| [19] | LU Z J, ZHU L J, LIANG G B,et al.MORE FLORET1 controls anther development by negatively regulating key tapetal genes in both diploid and tetraploid rice[J].Plant Physiology,2024,195(3):1981-1994. |
| [20] | CAO Y P, LI K, LI Y L,et al.MYB transcription factors as regulators of secondary metabolism in plants[J].Biology,2020,9(3):61. |
| [21] | WANG S S, JIANG R Y, FENG J,et al.Overexpression of transcription factor FaMYB63 enhances salt tolerance by directly binding to the SOS1 promoter in Arabidopsis thaliana [J].Plant Molecular Biology,2024,114(2):32. |
| [22] | PAZ-ARES J, GHOSAL D, WIENAND U,et al.The regulatory c1 locus of Zea mays encodes a protein with homology to myb proto‐oncogene products and with structural similarities to transcriptional activators[J].The EMBO Journal,1987,6(12):3553-3558. |
| [23] | DELUC L, BARRIEU F, MARCHIVE C,et al.Characterization of a grapevine R2R3-MYB transcription factor that regulates the phenylpropanoid pathway[J].Plant Ph-ysiology,2006,140(2):499-511. |
| [24] | VIMOLMANGKANG S, HAN Y P, WEI G C,et al.An apple MYB transcription factor,MdMYB3,is involved in regulation of anthocyanin biosynthesis and flower development[J].BMC Plant Biology,2013,13:176. |
| [25] | LI S H, DONG Y X, LI D L,et al.Eggplant transcription factor SmMYB5 integrates jasmonate and light signaling during anthocyanin biosynthesis[J].Plant Physiology,2024,194(2):1139-1165. |
| [26] | ZHOU L J, GENG Z Q, WANG Y X,et al.A novel transcription factor CmMYB012 inhibits flavone and anthocyanin biosynthesis in response to high temperatures in chrysanthemum[J].Horticulture Research,2021,8(1):248. |
| [27] | WANG X X, WANG J C, LIU Z Y,et al.The R2R3 MYB gene TaMYB305 positively regulates anther and pollen development in thermo-sensitive male-sterility wheat with Aegilops kotschyi cytoplasm[J].Planta,2024,259(3):64. |
| [28] | WANG C H, YAO H X, WANG C,et al.Transcription factor CsMYB36 regulates fruit neck length via mediating cell expansion in cucumber[J].Plant Physiology,2024,195(2):958-969. |
| [29] | ZHU L, GUAN Y X, ZHANG Z H,et al. CmMYB8 encodes an R2R3 MYB transcription factor which represses lignin and flavonoid synthesis in chrysanthemum[J].Plant Physiology and Biochemistry,2020,149:217-224. |
| [30] | SUN C D, HUANG H Z, XU C J,et al.Biological activities of extracts from Chinese bayberry (Myrica rubra Sieb.et Zucc.):a review[J].Plant Foods for Human Nutrition,2013,68(2):97-106. |
| [31] | 庞沁,陈成标,邹登朗,等.黑果枸杞花青素研究概述[J].农家参谋,2022(18):114-116. |
| PANG Q, CHEN C B, ZOU D L,et al.Overview of anthocyanin research in Lycium ruthenicum [J].The Farmers Consultant,2022(18):114-116. | |
| [32] | WAN S Z, LI C F, MA X D,et al.PtrMYB57 contributes to the negative regulation of anthocyanin and proanthocyanidin biosynthesis in poplar[J].Plant Cell Reports,2017,36(8):1263-1276. |
| [33] | SHI L Y, CHEN X, WANG K,et al.MrMYB6 from Chinese bayberry (Myrica rubra) negatively regulates anthocyanin and proanthocyanidin accumulation[J].Frontiers in Plant Science,2021,12:685654. |
| [34] | WANG S, ZHANG Z, LI L X,et al.Apple MdMYB306-like inhibits anthocyanin synthesis by directly interacting with MdMYB17 and MdbHLH33[J].The Plant Journal,2022,110(4):1021-1034. |
| [35] | ZHANG F, WANG J X, DING T T,et al.MYB2 and MYB108 regulate lateral root development by interacting with LBD29 in Arabidopsis thaliana [J].Journal of Integrative Plant Biology,2024,66(8):1675-1687. |
| [36] | JIANG L J, ZHANG D H, LIU C,et al.MdGH3.6 is targeted by MdMYB94 and plays a negative role in apple water-deficit stress tolerance[J].The Plant Journal,2022,109(5):1271-1289. |
| [37] | HAN Y F, QU M H, LIU Z C,et al.Transcription factor FveMYB117a inhibits axillary bud outgrowth by regulating cytokinin homeostasis in woodland strawberry[J].The Plant Cell,2024,36(6):2427-2446. |
| [38] | JIANG P F, LIN X Y, BIAN X Y,et al.Ectopic expression of Populus MYB10 promotes secondary cell wall thickening and inhibits anthocyanin accumulation[J].Plant Physiology and Biochemistry,2022,172:24-32. |
| [1] | Nan LI, Xiaoxia TIAN, Peichun MAO, Mingli ZHENG, LIN MENG, Lan YUN. Phenotype and Pigment Analysis of Flower Organs of Iris lactea var. chinensis [J]. Bulletin of Botanical Research, 2024, 44(1): 51-61. |
| [2] | Qinghua YANG, Binjie GE, Dasheng ZHANG. Observation on the Formation and Composition Analysis of Blue Pollen of Caryopterisincana [J]. Bulletin of Botanical Research, 2023, 43(5): 794-800. |
| [3] | Zhezhe LI, Yidan ZHANG, Bo WANG, Zhenghao WANG, Lu WANG, Beibei CHENG. Changes and Influencing Factors of Anthocyanins in Hibiscus syriacus During Flowering [J]. Bulletin of Botanical Research, 2023, 43(4): 550-561. |
| [4] | Chen-Chen ZHOU, Yi-Liang LI, Zhe WANG, Fen-Cheng ZHAO, Sui-Ying ZHONG, Chang-Ming LIN, Zhi-Qiang TAN, Fu-Ming LI, Hui-Shan WU, Wen-Bing GUO, Fang-Yan LIAO, Jian-Ge WANG. Screening and Expression Analysis of Different Genes for Oleoresin Production in the Specific Period of Pinus elliottii×P.caribaea by RNA-Seq Technology [J]. Bulletin of Botanical Research, 2021, 41(3): 419-428. |
| [5] | Li-Qiang ZHAO, Chun-Miao SHAN, Sheng-Xiang ZHANG, Yuan-Yuan SHI, Ke-Long MA, Jia-Wen WU. Identification of Key Enzyme Genes Involved in Anthocyanin Synthesis pathway in Clinopodium gracile by Transcriptome Analysis [J]. Bulletin of Botanical Research, 2020, 40(6): 886-896. |
| [6] | LIANG Ling, JIANG Jie-Bei, ZHANG Teng-Ju, Lü Jia-Yao, CHEN Xiao-Hong. Physiological Characteristics of Davidia involucrata Bracts and Leaves with Different Colors [J]. Bulletin of Botanical Research, 2020, 40(4): 505-513. |
| [7] | YANG Yun, CHEN Meng-Jiao, DU Qing-Xin, ZHU Jing-Le, DU Hong-Yan, YANG Shao-Bin. SNP Sites Developed by Whole Genome Resequencing Analysis in Eucommia ulmoides ‘Hongye’ [J]. Bulletin of Botanical Research, 2019, 39(6): 947-954. |
| [8] | QIN Lin-Lin, ZHANG Xi, JIANG Cheng, LI Li. Cloning and Functional Analysis of BpZFP4 Promoter from Birch(Betula platyphylla) [J]. Bulletin of Botanical Research, 2019, 39(6): 917-926. |
| [9] | LIU Xiong-Fang, LI Tai-Qiang, ZHANG Xu, LI Zheng-Hong, WAN You-Ming, AN Jing, LIU Xiu-Xian, MA Hong. Characteristic Analysis of Microsatellites in the Transcriptome of Phyllanthus emblica,an Important Edible and Medicinal Plant [J]. Bulletin of Botanical Research, 2019, 39(2): 294-302. |
| [10] | SHAO Wan-Lu, LI Yue-Ling, GAO Song, LI Jun-Min, LIANG Zong-Suo. Effects of Light Intensity on the Fruit Coloration and Anthocyanian Biosynthesis in Fragaria×ananassa Duch. ‘Benihoppe’ and the Possible Molecular Mechanism [J]. Bulletin of Botanical Research, 2018, 38(5): 661-668. |
| [11] | YU Ting-Ting, NI Xiu-Zhen, GAO Li-Hong, HAN Guo-Jun, ZHU Chang-Fu, SHENG Yan-Min. Advances in Study of Dihydroflavonol 4-Reductase(DFR) Genes of Higher Plants [J]. Bulletin of Botanical Research, 2018, 38(4): 632-640. |
| [12] | XU Meng-Ke, LI Dan, MENG Lai-Sheng, JIANG Ji-Hong. Arabidopsis thaliana Transcription Factor Ethylene-insensitive3(EIN3) Inhibits the Synthesis of Anthocyanins [J]. Bulletin of Botanical Research, 2018, 38(1): 148-154. |
| [13] | XU Yi-Qian1;YUAN Yuan1;TAO Xiu-Hua2;YANG Juan3;SHI Yi-Min1;TANG Dong-Qin1*. Main Anthocyanin Profiles in Petals of Freesia hybrida [J]. Bulletin of Botanical Research, 2016, 36(2): 184-189. |
| [14] | XU Zhi-Ru1,2,3;MA Jing1,2,3;CUI Guo-Xin1,2,3;LIU Tong1,2,3;LIU Guan-Jun1,2,3. Functional Identification and Promoter Preliminary Analysis of Flavonoid 3′-Hydroxylase Genes in Turnip [J]. Bulletin of Botanical Research, 2015, 35(4): 572-582. |
| [15] | LI Ya-Li;HU Zong-Li;ZHANG Bin;ZHU Ming-Ku;CHEN Guo-Ping*. Effect of Light on Anthocyanin Biosynthesis and Metabolism of Brassica oleracea L. and Analysis on Expression Pattern of Anthocyanin Structural Genes [J]. Bulletin of Botanical Research, 2012, 32(4): 397-401. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||