Bulletin of Botanical Research ›› 2022, Vol. 42 ›› Issue (5): 821-829.doi: 10.7525/j.issn.1673-5102.2022.05.013
• Molecular biology • Previous Articles Next Articles
Denggao LI, Rui LIN, Qinghui MU, Na ZHOU, Yanru ZHANG, Wei BAI( )
)
Received:2022-03-15
Online:2022-09-20
Published:2022-09-15
Contact:
Wei BAI
E-mail:baiwei@imau.edu.cn
About author:LI Denggao(1990—),male,doctor,mainly engaged in research of molecular biology of plant stress resistance.
Supported by:CLC Number:
Denggao LI, Rui LIN, Qinghui MU, Na ZHOU, Yanru ZHANG, Wei BAI. Cloning and Functional Analysis of StNPR4 gene in Solanum tuberosum[J]. Bulletin of Botanical Research, 2022, 42(5): 821-829.
Add to citation manager EndNote|Ris|BibTeX
URL: https://bbr.nefu.edu.cn/EN/10.7525/j.issn.1673-5102.2022.05.013
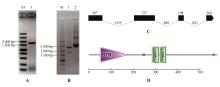

Fig.2
PCR amplification,enzyme digestion and structure prediction of StNPR4A.PCR amplification of StNPR4 CDS(M.DNA Marker;1.StNPR4 CDSfragments);B.Enzyme digestion of StNPR4 cloning vector(M.DNA Marker; 1.Double digestion of StNPR4-pMD19-T vector;2.StNPR4-pMD19-T vector);C.StNPR4 gene structure;D.StNPR4 protein structural domains
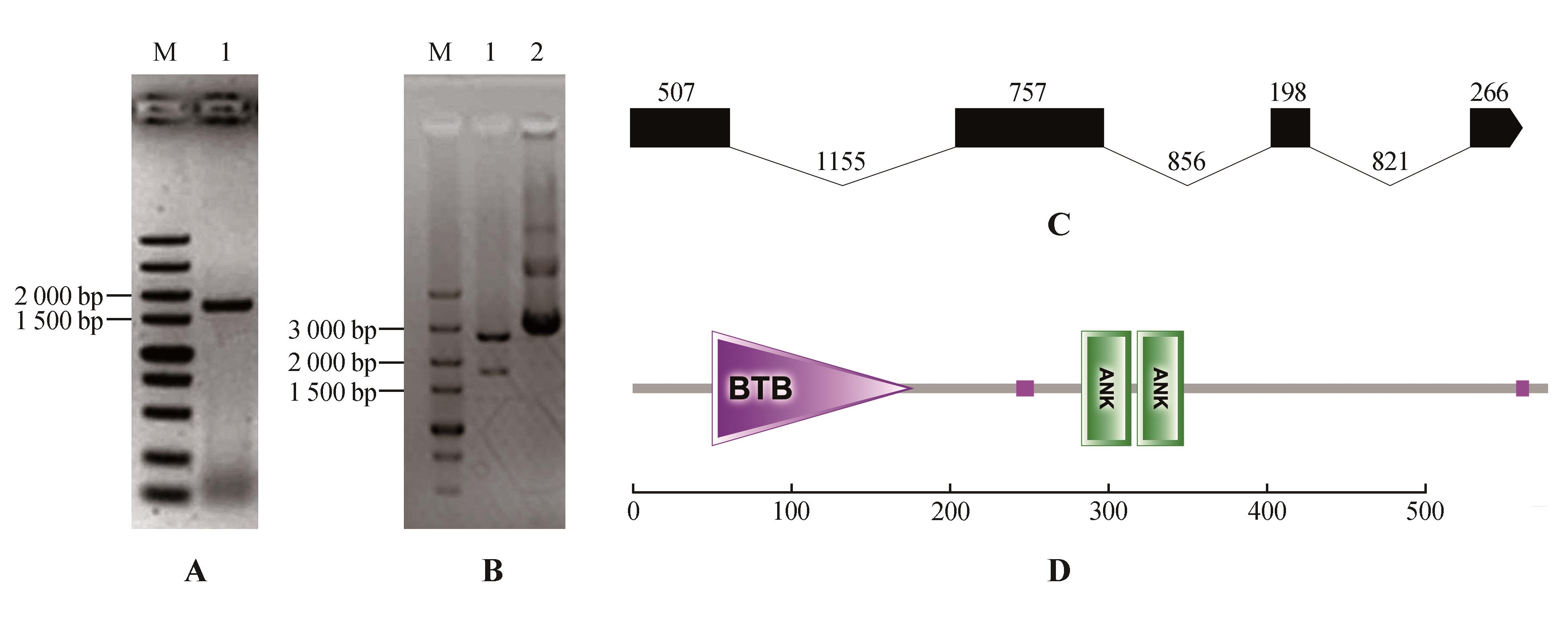
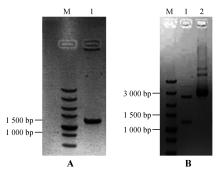
Fig.5
PCR amplification of StNPR4 promoter and enzyme digestion of the vectorA.PCR amplification of the StNPR4 promoter(M.DNA Marker;1.StPRONPR4 fragments at 1 246 bp;B.Enzyme digestion identification of StNPR4 cloning vector(M.DNA Marker;1.StPRONPR4 fragments at 1 246 bp;2.StproNPR4 -pENTR L16 vector)
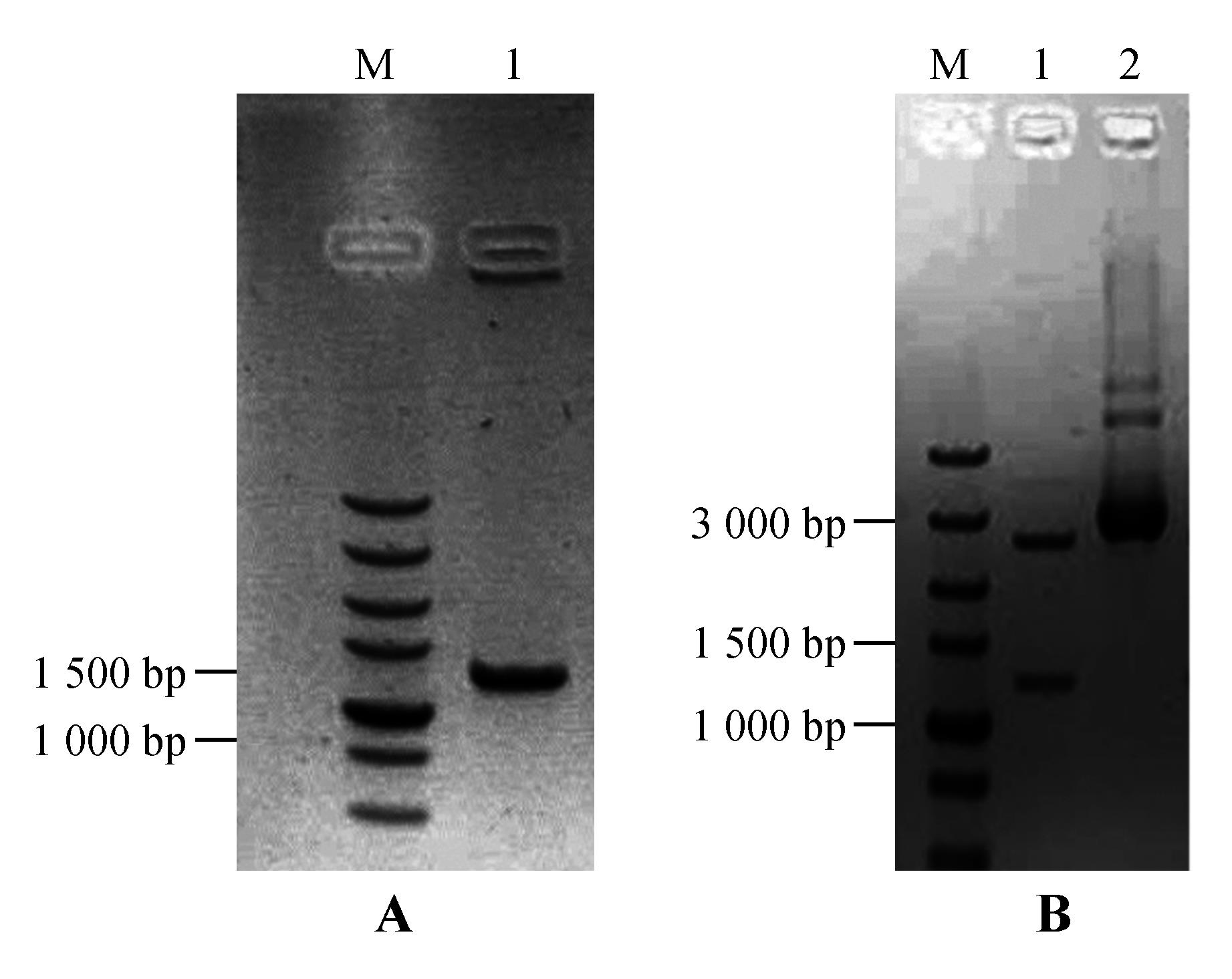
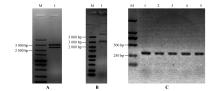
Fig.6
Enzyme digestion identification of vectors and PCR identification of transgenic plantsA.Identification of StPRONPR4-StNPR4 pENTR L16 cloning vector by enzymatic digestion;M.DNA Marker;1.StPRONPR4-StNPR4 fragments at 3 021 bp;B.Identification of StPRONPR4-StNPR4 pORE O4 binary expression vector by enzymatic digestion;M.DNA Marker;1.StPRONPR4-StNPR4 fragments at 3 021 bp;C.PCR identification of different StNPR4 transgenic lines;M.DNA Marker;1-5.Different StNPR4 transgenic lines
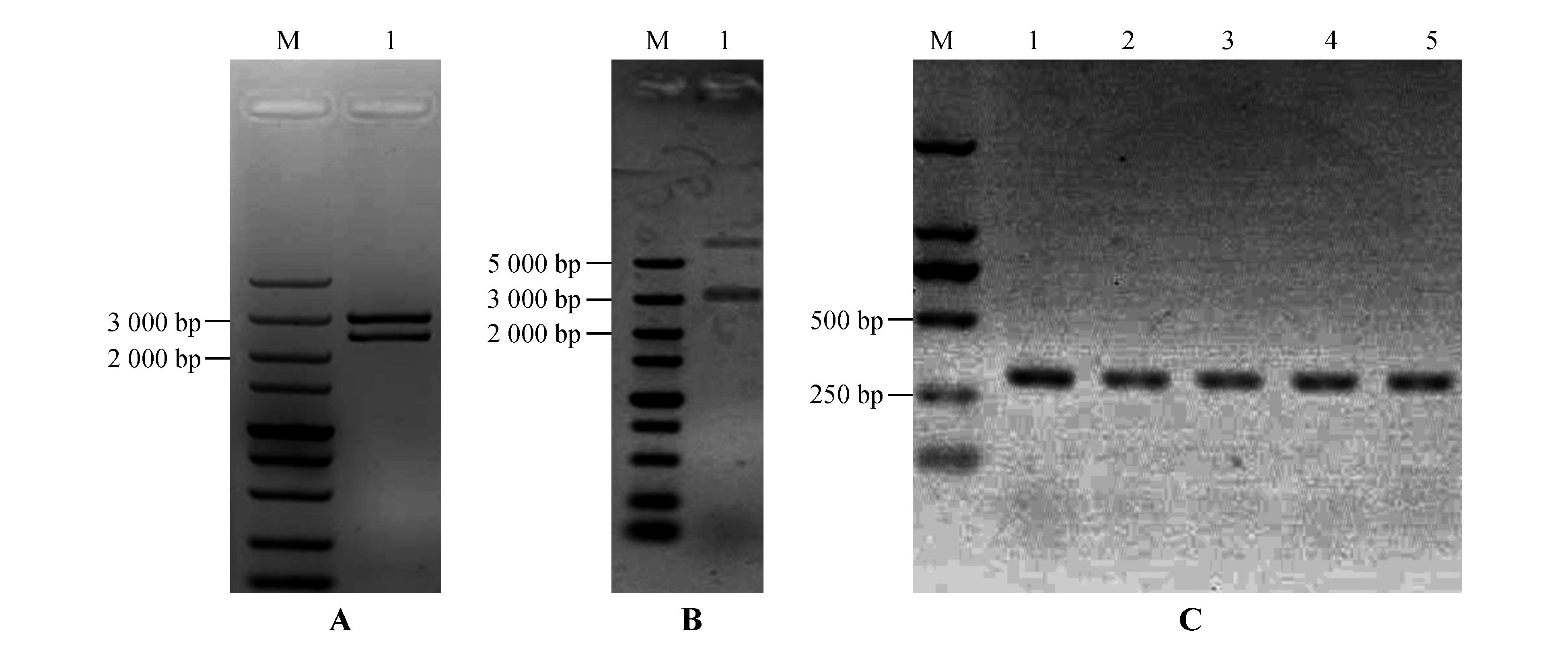
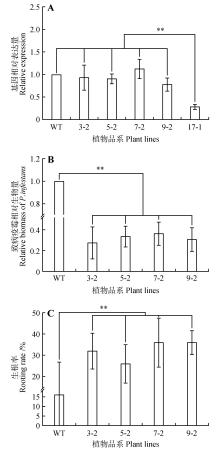
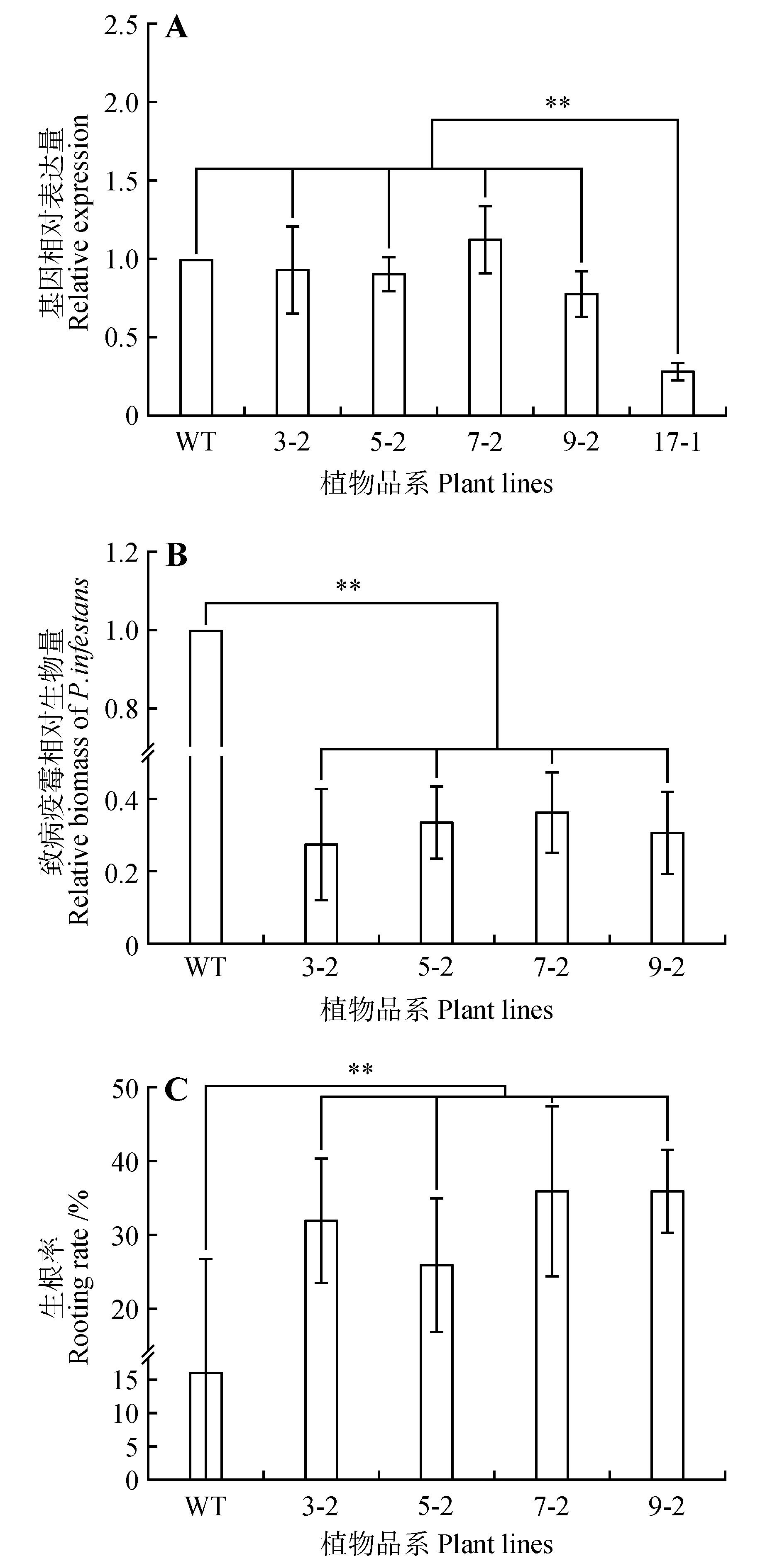
Fig.7
Functional analysis of StNPR4 transgenic plantsA.StNPR4 expression level in transgenic lines;B.Relative biomass of pathogens after inoculation of transgenic lines with phytophthora infestans;C.Rooting rates of transgenic lines after treatment with 150 mmol·L-1 NaCl;* represents P<0.05,** represents P<0.01,and equal variance t-test was used to analyze each experimental group and the control group at the same time point,the same as below
| 1 | CAO H, BOWLING S A, GORDON A S,et al.Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance[J].The Plant Cell,1994,6(11):1583-1592. |
| 2 | PIETERSE C M J, S C MVAN WEES, AVAN PELT J,et al.A novel signaling pathway controlling induced systemic resistance in Arabidopsis [J].The Plant Cell,1998,10(9):1571-1580. |
| 3 | SPOEL S H, KOORNNEEF A, CLAESSENS S M C,et al.NPR1 modulates cross-talk between salicylate-and jasmonate-dependent defense pathways through a novel function in the cytosol[J].The Plant Cell,2003,15(3):760-770. |
| 4 | MOU Z L, FAN W H, DONG X N.Inducers of plant systemic acquired resistance regulate NPR1 function through redox changes[J].Cell,2003,113(7):935-944. |
| 5 | ZHANG Y L, TESSARO M J, LASSNER M,et al.Knockout analysis of Arabidopsis transcription factors TGA2,TGA5,and TGA6 reveals their redundant and essential roles in systemic acquired resistance[J].The Plant Cell,2003,15(11):2647-2653. |
| 6 | JAYAKANNAN M, BOSE J, BABOURINA O,et al.Salicylic acid in plant salinity stress signalling and tolerance[J].Plant Growth Regulation,2015,76(1):25-40. |
| 7 | JAYAKANNAN M, BOSE J, BABOURINA O,et al.The NPR1-dependent salicylic acid signalling pathway is pivotal for enhanced salt and oxidative stress tolerance in Arabidopsis [J].Journal of Experimental Botany,2015,66(7):1865-1875. |
| 8 | SARISOY U, SECKIN DINLER B, TASCI E.The effects of NPR1 dependent salicylic acid change in increasing salt tolerance of soybean leaves by acclimation[J].Notulae Botanicae Horti Agrobotanici Cluj-Napoca,2018,46(2):356-364. |
| 9 | 刘晓东,李月,王若仲,等.过表达GH3-5提高拟南芥抗旱的分子机制[J].南京农业大学学报,2016,39(4):557-562. |
| LIU X D, LI Y, WANG R Z,et al.Molecular mechanism of drought tolerance conferred by overexpression of GH3-5[J].Journal of Nanjing Agricultural University,2016,39(4):557-562. | |
| 10 | OLATE E, JIMÉNEZ-GÓMEZ J M, HOLUIGUE L,et al.NPR1 mediates a novel regulatory pathway in cold acclimation by interacting with HSFA1 factors[J].Nature Plants,2018,4(10):811-823. |
| 11 | ZHOU M, WANG W, KARAPETYAN S,et al.Redox rhythm reinforces the circadian clock to gate immune response[J].Nature,2015,523(7561):472-476. |
| 12 | ZHANG Y L, CHENG Y T, QU N,et al.Negative regulation of defense responses in Arabidopsis by two NPR1 paralogs[J].The Plant Journal,2006,48(5):647-656. |
| 13 | DING Y L, SUN T J, AO K,et al.Opposite roles of salicylic acid receptors NPR1 and NPR3/NPR4 in transcriptional regulation of plant immunity[J].Cell,2018,173(6):1454-1467. |
| 14 | HEPWORTH S R, ZHANG Y L, MCKIM S,et al.BLADE-ON-PETIOLE-dependent signaling controls leaf and floral patterning in Arabidopsis [J].The Plant Cell,2005,17(5):1434-1448. |
| 15 | SPOEL S H, MOU Z L, TADA Y,et al.Proteasome - mediated turnover of the transcription coactivator NPR1 plays dual roles in regulating plant immunity[J].Cell,2009,137(5):860-872. |
| 16 | LIU Y N, SUN T J, SUN Y L,et al.Diverse roles of the salicylic acid receptors NPR1 and NPR3/NPR4 in plant immunity[J].The Plant Cell,2020,32(12):4002-4016. |
| 17 | BACKER R, NAIDOO S,BERG NVAN DEN.The NONEXPRESSOR OF PATHOGENESIS-RELATED GENES1(NPR1) and related family:mechanistic insights in plant disease resistance[J].Frontiers in Plant Science,2019,10:102. |
| 18 | 王攀,蔡兆明,廖静静,等.大白菜NPR家族基因鉴定及表达模式分析[J].分子植物育种,2021,19(3):746-758. |
| WANG P, CAI Z M, LIAO J J,et al.Genome-wide identification and expression pattern analysis of histidine NPR family genes in Brassica rapa ssp. pekinensis [J].Molecular Plant Breeding,2021,19(3):746-758. | |
| 19 | 李雅倩,王林,宋婧含,等.小麦NPR1基因家族鉴定及其表达分析[J].南方农业学报,2021,52(9):2339-2349. |
| LI Y Q, WANG L, SONG J H,et al.Genome-wide identification and expression analysis of wheat NPR1 gene family[J].Journal of Southern Agriculture,2021,52(9):2339-2349. | |
| 20 | 江慎秀,尚海,李涛,等.油桐NPR1家族全基因组鉴定及表达模式分析[J].植物遗传资源学报,2021,22(2):521-531. |
| JIANG S X, SHANG H, LI T,et al.Genome-wide identification and expression pattern analysis of NPR1 family in Vernicia fordii [J].Journal of Plant Genetic Resources,2021,22(2):521-531. | |
| 21 | ZHANG J K, JIAO P, ZHANG C,et al.Apple NPR1 homologs and their alternative splicing forms may contribute to SA and disease responses[J].Tree Genetics & Genomes,2016,12(5):92. |
| 22 | 淳俊,桑有顺,冯焱,等.四川省马铃薯晚疫病研究进展[J].中国农学通报,2018,34(30):136-139. |
| CHUN J, SANG Y S, FENG Y,et al.Potato late blight in Sichuan:research progress[J].Chinese Agricultural Science Bulletin,2018,34(30):136-139. | |
| 23 | TIAN L X, YANG J Q, HOU W J,et al.Molecular cloning and characterization of a P-glycoprotein from the diamondback moth,Plutella xylostella(Lepidoptera:Plutellidae) [J].International Journal of Molecular Sciences,2013,14(11):22891-22905. |
| 24 | WITHERS J, DONG X N.Posttranslational modifications of NPR1:a single protein playing multiple roles in plant immunity and physiology[J].PLoS Pathogens,2016,12(8):e1005707. |
| 25 | BAI W, CHERN M, RUAN D L,et al.Enhanced disease resistance and hypersensitivity to BTH by introduction of an NH1/OsNPR1 paralog[J].Plant Biotechnology Journal,2011,9(2):205-215. |
| 26 | ZAVALIEV R, MOHAN R, CHEN T Y,et al.Formation of NPR1 condensates promotes cell survival during the plant immune response[J].Cell,2020,182(5):1093-1108. |
| 27 | SEO S Y, WI S J, PARK K Y.Functional switching of NPR1 between chloroplast and nucleus for adaptive response to salt stress[J].Scientific Reports,2020,10(1):4339. |
| 28 | DIVI U K, RAHMAN T, KRISHNA P.Brassinosteroid - mediated stress tolerance in Arabidopsis shows interactions with abscisic acid,ethylene and salicylic acid pathways[J].BMC Plant Biology,2010,10:151. |
| 29 | QUILIS J, PEÑAS G, MESSEGUER J,et al.The Arabidopsis AtNPR1 inversely modulates defense responses against fungal,bacterial,or viral pathogens while conferring hypersensitivity to abiotic stresses in transgenic rice[J].Molecular Plant-Microbe Interactions,2008,21(9):1215-1231. |
| 30 | HAO L, ZHAO Y, JIN D D,et al.Salicylic acid - altering Arabidopsis mutants response to salt stress[J].Plant and Soil,2012,354(1/2):81-95. |
| 31 | XU Y Q, WANG H, QIN R L,et al.Characterization of NPR1 and NPR4 genes from mulberry(Morus multicaulis) and their roles in development and stress resistance[J].Physiologia Plantarum,2019,167(3):302-316. |
| [1] | Lipeng ZHANG, Mei WU, Hongpeng WANG, Tianyu LI. Salt Stress-related RcNAC22 Gene Cloning and Function Analysis in Rhodiolacrenulata [J]. Bulletin of Botanical Research, 2026, 46(2): 246-258. |
| [2] | Haiyan CAO, Kaiwen TIAN, Xiaoyu JIA, Xuefeng HAO, Zhuping JIN. The Role of AtMST1 in Regulating Salt Tolerance via H2S Synthesis in Arabidopsis Revealed by CRISPR/Cas9 Knockout [J]. Bulletin of Botanical Research, 2026, 46(2): 259-269. |
| [3] | Jingjing ZHANG, Haixia LUO, Yanping HONG, Shicheng ZHAO. Optimization of Agrobacterium rhizogenes-Mediated Genetic Transformation System of Solanum nigrum [J]. Bulletin of Botanical Research, 2026, 46(1): 111-119. |
| [4] | Yuping LIU, Mingjun YU, Yu ZHANG, Xu SU, Xiaoli LI, Qian YANG, Xueli LIU. Cloning and Drought Tolerance Functional Validation of PvAQP Gene in Psammochloa villosa (Poaceae) [J]. Bulletin of Botanical Research, 2024, 44(6): 914-925. |
| [5] | Xiuying MA, Jinke LI, Xiaoyang ZHOU, Shaoliang CHEN. Research Progress of Ca2+-ATPase Involved in Regulation of Plant Salt Tolerance [J]. Bulletin of Botanical Research, 2024, 44(5): 641-654. |
| [6] | Jiayu CAO, Lina CAO, Qiaoyi ZHANG, Shuang ZHANG, Zhibao HU, Tangrui ZHAO, Zhiru XU, Chunming LI, Xiankui QUAN, Guanjun LIU. Cloning and Expression Activity Analysis of Rubisco Small Subunit Gene Promoter in Populus × xiaohei [J]. Bulletin of Botanical Research, 2024, 44(4): 625-633. |
| [7] | Xiaoqian WU, Xu HE, Jinghui GAO, Shuang LI. Germplasm Innovation and Characteristic Analysis of Transgenic PsnNAC007Populus simonii×P. nigra with High Drought Tolerance [J]. Bulletin of Botanical Research, 2024, 44(3): 349-360. |
| [8] | Dongyue WANG, Ruyue WANG, Maotong SUN, Cuishuang LIU, Jihong LI. Establishment of Genetic Transformation System for ‘Populus leucopyramidalis 1’ and Transformation of Insect Resistance Gene [J]. Bulletin of Botanical Research, 2024, 44(3): 361-369. |
| [9] | Hengfeng ZHANG, Yangwu HE, Huanchao ZHANG, Qingcui WEI. Metabolomics Analysis of Lagerstroemia indica in Response to Salt and Alkali Stress [J]. Bulletin of Botanical Research, 2024, 44(3): 420-430. |
| [10] | Fazhi FANG, Huiying GUI, Zhaojia LI, Xiaofeng ZHANG. Physiological Adaptation of Six Mangrove Seedlings to Different Salinity [J]. Bulletin of Botanical Research, 2023, 43(6): 881-889. |
| [11] | Xu HE, Yuan GAO, Qunye ZHANG, Chenguang ZHOU, Wei LI, Shuang LI. Establishment and Application of Genetic Transformation System for Populus simonii×P. nigra ‘Baicheng’ [J]. Bulletin of Botanical Research, 2023, 43(5): 667-678. |
| [12] | Lei XU, Xiao XU, Qinsong LIU. Effects of Exogenous Salicylic Acid on Antioxidant System and Gene Expression of Davidia involucrata Seedlings under Salt Stress [J]. Bulletin of Botanical Research, 2023, 43(4): 572-581. |
| [13] | Haonan ZHANG, Shanshan CHEN, Jianmin XU, Ping LUO, Xiaoping WANG, Zhiru XU, Chunjie FAN. Cloning and Functional Analysis of EgrWAT1 Gene in Eucalyptus grandis [J]. Bulletin of Botanical Research, 2023, 43(4): 601-611. |
| [14] | Li LI, Xin WANG, Yuejing ZHANG, Lingyun JIA, Hailong PANG, Hanqing FENG. Effects of Abiotic Stresses on the Intracellular and Extracellular ATP Levels of Tobacco Suspension Cells [J]. Bulletin of Botanical Research, 2023, 43(2): 179-185. |
| [15] | Shixian LIAO, Yuting WANG, Liben DONG, Yongmei GU, Fenglin JIA, Tingbo JIANG, Boru ZHOU. Function Analysis of the Transcription Factor PsnbZIP1 of Populus simonii×P. nigra in Response to Salt Stress [J]. Bulletin of Botanical Research, 2023, 43(2): 288-299. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||