Bulletin of Botanical Research ›› 2021, Vol. 41 ›› Issue (5): 729-737.doi: 10.7525/j.issn.1673-5102.2021.05.011
Previous Articles Next Articles
Tian-Tian DU1, Hui-Ping LI1, Bo-Ya WANG1, Qian OU1, Yan HUANG1, Ying CAO1, Shang-Lian HU1,2( )
)
Received:2020-12-29
Online:2021-09-20
Published:2021-07-05
Contact:
Shang-Lian HU
E-mail:hushanglian@126.com;hashanglian@126.com
About author:DU Tian-Tian(1995—),female,master,focused on biochemistry and molecular biology.
Supported by:CLC Number:
Tian-Tian DU, Hui-Ping LI, Bo-Ya WANG, Qian OU, Yan HUANG, Ying CAO, Shang-Lian HU. Cloning and Promotor Analysis of DfMYB3 from Dendrocalamus farinosus[J]. Bulletin of Botanical Research, 2021, 41(5): 729-737.
Add to citation manager EndNote|Ris|BibTeX
URL: https://bbr.nefu.edu.cn/EN/10.7525/j.issn.1673-5102.2021.05.011
Table 1
Primer sequences
引物名称 Primer name | 引物序列(5′→3′) Primer sequences(5′→3′) |
|---|---|
| DfMYB3-F | ATGGGGAGGCATTCTTGTTG |
| DfMYB3-R | CTAGATATGCTCAAAAGATA |
| RT-DfMYB3-F | AAGCAACGGTATGGTATTCGAGCCA |
| RT-DfMYB3-R | GCACATATCCCTTCCATGTTGAATT |
| Tublin-F | GCCGTGAATCTCATCCCCTT |
| Tublin-R | TTGTTCTTGGCATCCCACAT |
| DfMYB3-F(KpnⅠ) | GGGGTACCCCATGGGGAGGCATTCTTGTTG |
| DfMYB3-R(HindⅢ) | CCCAAGCTTGGGGATATGCTCAAAAGATA |
| P3 | TTGCTTGTAGCAACAAGAATGC |
| P2 | TGTTCCCAAGGACAGCATGGAG |
| P1 | TTTCAGGGCGATTCTTATCATA |
| pDfMYB3-F(HindⅢ) | CCAAGCTTGGTATCGAGTTTCGTGTTCCCAAG |
| pDfMYB3-R(XbaI) | GCTCTAGAGCTTCGGTGGAATGCACAACCTGA |

Fig.1
Cloning of DfMYB3 gene from Dendrocalamus farinosus and bioinformatics analysisA.Cloning of DfMYB3 gene from Dendrocalamus farinosus;B.Conserved domain analysis of DfMYB3;C.Secondary structure analysis of DfMYB3(Blue.Alpha helix;Purple.Random coil;Red.Extended strand;Green.Beta turn);D.3D structure analysis of DfMYB3
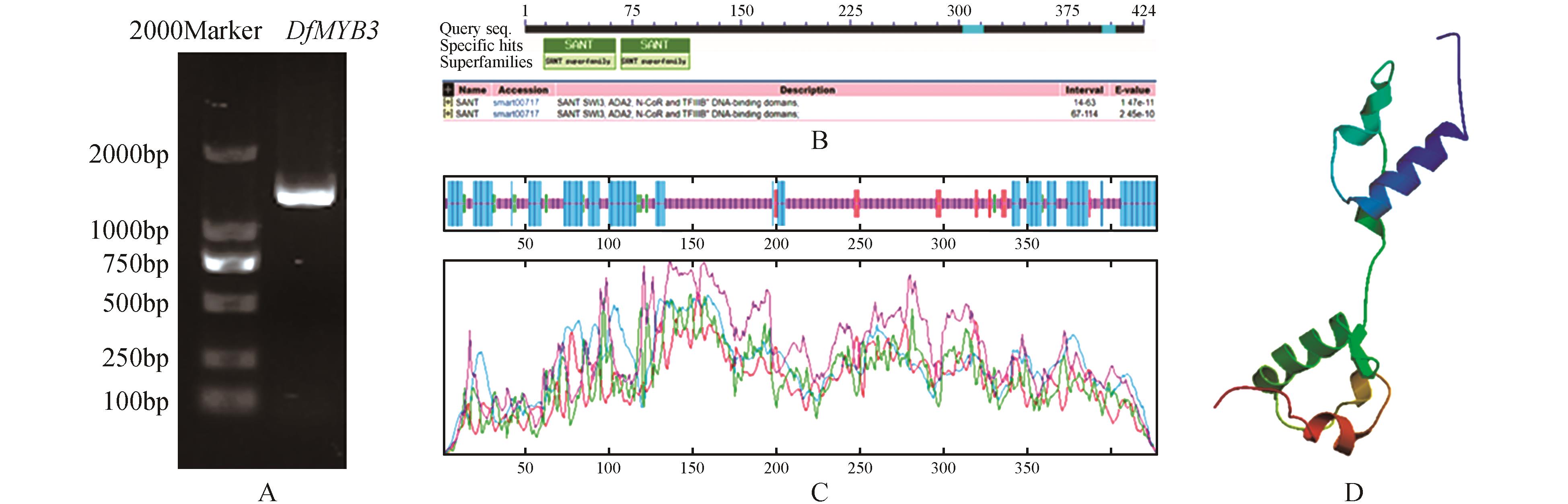
Table 2
Element analysis of DfMYB3 promoter
元件 Site name | 物种 Organism | 位置 Position | 链 Strand | 评分 Matrix score | 序列 Sequence | 功能 Function |
|---|---|---|---|---|---|---|
| ABRE | 拟南芥Arabidopsis thaliana | 705 | + | 5 | ACGTG | cis-acting element involved in the abscisic acid responsiveness |
| ABRE | 大麦Hordeum vulgare | 976 | _ | 9 | CGTACGTGCA | cis-acting element involved in the abscisic acid responsiveness |
| AuxRR-core | 烟草Nicotiana tabacum | 537 | - | 7 | GGTCCAT | cis-acting regulatory element involved in auxin responsiveness |
| Box 4 | 欧芹Petroselinum crispum | 96 | + | 6 | ATTAAT | part of a conserved DNA module involved in light responsiveness |
| CGTCA-motif | 大麦Hordeum vulgare | 191 | + | 5 | CGTCA | cis-acting regulatory element involved in the MeJA-responsiveness |
| CGTCA-motif | 大麦Hordeum vulgare | 813 | - | 5 | CGTCA | cis-acting regulatory element involved in the MeJA-responsiveness |
| CGTCA-motif | 大麦Hordeum vulgare | 604 | - | 5 | CGTCA | cis-acting regulatory element involved in the MeJA-responsiveness |
| CGTCA-motif | 大麦Hordeum vulgare | 854 | - | 5 | CGTCA | cis-acting regulatory element involved in the MeJA-responsiveness |
| G-Box | 豌豆Pisum sativum | 704 | - | 6 | CACGTT | cis-acting regulatory element involved in light responsiveness |
| GARE-motif | 甘草Brassica oleracea | 1017 | - | 7 | TCTGTTG | gibberellin-responsive element |
| GATA-motif | 拟南芥Arabidopsis thaliana | 1574 | - | 7 | GATAGGA | part of a light responsive element |
| GT1-motif | 拟南芥Arabidopsis thaliana | 673 | - | 6 | GGTTAA | light responsive elemen |
| MBS | 拟南芥Arabidopsis thaliana | 841 | + | 6 | CAACTG | MYB binding site involved in drought-inducibility |
| MBS | 拟南芥Arabidopsis thaliana | 1045 | + | 6 | CAACTG | MYB binding site involved in drought-inducibility |
| P-box | 水稻Oryza sativa | 775 | - | 7 | CCTTTTG | gibberellin-responsive element |
| TCA-element | 烟草Nicotiana tabacum | 1108 | - | 9 | CCATCTTTTT | cis-acting element involved in salicylic acid responsiveness |
| TGACG-motif | 大麦Hordeum vulgare | 191 | - | 5 | TGACG | cis-acting regulatory element involved in the MeJA-responsiveness |
| TGACG-motif | Hordeum vulgare | 813 | + | 5 | TGACG | cis-acting regulatory element involved in the MeJA-responsiveness |
| TGACG-motif | Hordeum vulgare | 604 | + | 5 | TGACG | cis-acting regulatory element involved in the MeJA-responsiveness |
| TGACG-motif | Hordeum vulgare | 854 | + | 5 | TGACG | cis-acting regulatory element involved in the MeJA-responsiveness |
| 1 | Pazares J,Ghosal D,Wienand U,et al.The regulatory c1 locus of Zea mays encodes a protein with homology to myb proto-oncogene products and with structural similarities to transcriptional activators [J].Embo Journal,1987,6(12): 3553-3558. |
| 2 | Loguercio L L,Zhang J Q,Wilkins T A.Differential regulation of six novel MYB-domain genes defines two distinct expression patterns in allotetraploid cotton(Gossypium hirsutum L.) [J].Molecular and General Genetics MGG,1999,261(4-5):660-671. |
| 3 | Miyake K,Ito T,Senda M,et al.Isolation of a subfamily of genes for R2R3-MYB transcription factors showing up-regulated expression under nitrogen nutrient-limited conditions[J].Plant Molecular Biology,2003,53(1-2):237-245. |
| 4 | Chen Y H,Yang X Y,He K,et al.The MYB transcription factor superfamily of Arabidopsis:expression analysis and phylogenetic comparison with the rice MYB family[J].Plant Molecular Biology,2006,60(1):107-124. |
| 5 | Stracke R,Werber M,Weisshaar B.The R2R3-MYB gene family in Arabidopsis thaliana[J].Current Opinion in Plant Biology,2001,4(5):447-456. |
| 6 | Suo J F,Liang X,Pu L,et al.Identification of GhMYB109 encoding a R2R3 MYB transcription factor that expressed specifically in fiber initials and elongating fibers of cotton(Gossypium hirsutum L.)[J].Biochimica et Biophysica Acta(BBA)-Gene Structure and Expression,2003,1630(1):25-34. |
| 7 | Kim W C,Kim J Y,Ko J H,et al.Identification of direct targets of transcription factor MYB46 provides insights into the transcriptional regulation of secondary wall biosynthesis[J].Plant Molecular Biology,2014,85(6):589-599. |
| 8 | Uimari A,Strommer J.Myb26:a MYB-like protein of pea flowers with affinity for promoters of phenylpropanoid genes[J].The Plant Journal,1997,12(6):1273-1284. |
| 9 | Ambawat S,Sharma P,Yadav N R,et al.MYB transcription factor genes as regulators for plant responses:an overview[J].Physiology and Molecular Biology of Plants,2013,19(3):307-321. |
| 10 | Huang J F,Chen F,Wu S Y,et al.Cotton GhMYB7 is predominantly expressed in developing fibers and regulates secondary cell wall biosynthesis in transgenic Arabidopsis[J].Science China Life Sciences,2016,59(2):194-205. |
| 11 | Park J S,Kim J B,Cho K J,et al.Arabidopsis R2R3-MYB transcription factor AtMYB60 functions as a transcriptional repressor of anthocyanin biosynthesis in lettuce(Lactuca sativa)[J].Plant Cell Reports,2008,27(6):985-994. |
| 12 | Li H L,Guo D,Peng S Q.Molecular and functional characterization of the JcMYB1,encoding a putative R2R3-MYB transcription factor in Jatropha curcas[J].Plant Growth Regulation,2015,75(1):45-53. |
| 13 | Wang W N,Sun Q,Cai C W,et al.Molecular characterization and alternative splicing of a MYB transcription factor gene in tumourous stem mustard and its response to abiotic stresses[J].Biologia,2016,71(5):538-546. |
| 14 | Fang W B,Ding W W,Zhao X L,et al.Expression profile and function characterization of the MYB type transcription factor genes in wheat(Triticum aestivum L.) under phosphorus deprivation[J].Acta Physiologiae Plantarum,2016,38(1):5. |
| 15 | Yao L M,Jiang Y N,Lu X X,et al.A R2R3-MYB transcription factor from Lablab purpureus induced by drought increases tolerance to abiotic stress in Arabidopsis[J].Molecular Biology Reports,2016,43(10):1089-1100. |
| 16 | Chen Y H,Cao Y Y,Wang L J,et al.Identification of MYB transcription factor genes and their expression during abiotic stresses in maize[J].Biologia Plantarum,2018,62(2):222-230. |
| 17 | 江泽慧.世界竹藤[M].沈阳:辽宁科学技术出版社,2002:367. |
| Jiang Z H.Bamboo and rattan in the world[M].Shenyang:Liaoning Science and Technology Press,2002:367. | |
| 18 | 郭晋隆,李国印,苏亚春,等.甘蔗R2R3-MYB类似基因Sc2RMyb1的克隆及表达特性分析[J].农业生物技术学报,2012,20(9):1009-1017. |
| Guo J L,Li G Y,Su Y C,et al.Cloning and expression analysis of a R2R3-MYB-like gene Sc2RMyb1 from sugarcane(Saccharum complex)[J].Journal of Agricultural Biotechnology,2012,20(9):1009-1017. | |
| 19 | Tian Q Y,Wang X Q,Li C F,et al.Functional characterization of the poplar R2R3-MYB transcription factor PtoMYB216 involved in the regulation of lignin biosynthesis during wood formation[J].PLoS One,2013,8(10):e76369. |
| 20 | Park M Y,Kang J Y,Kim S Y.Overexpression of AtMYB52 confers ABA hypersensitivity and drought tolerance[J].Molecules and Cells,2011,31(5):447-454. |
| 21 | Qin Y X,Wang M C,Tian Y C,et al.Over-expression of TaMYB33 encoding a novel wheat MYB transcription factor increases salt and drought tolerance in Arabidopsis[J].Molecular Biology Reports,2012,39(6):7183-7192. |
| 22 | Zhao K M,Bartley L E.Comparative genomic analysis of the R2R3 MYB secondary cell wall regulators of Arabidopsis,poplar,rice,maize,and switchgrass[J].BMC Plant Biology,2014,14:135. |
| 23 | 刘慧春,马广莹,朱开元,等.牡丹PsDREB转录因子基因的克隆及亚细胞定位[J].分子植物育种,2015,13(10):2290-2298. |
| Liu H C,Ma G Y,Zhu K Y,et al.Clonging and subcellular location of a DREB transcription gene of Paeonia suffruticosa[J].Molecular Plant Breeding,2015,13(10):2290-2298. | |
| 24 | Arenhart R A,Schunemann M,Neto L B,et al.Rice ASR1 and ASR5 are complementary transcription factors regulating aluminium responsive genes[J].Plant,Cell & Environment,2016,39(3):645-651. |
| 25 | 张顺仓,冯思念,顾雯,等.丹参中转录因子SmMYB87基因的克隆、亚细胞定位和表达分析[J].中草药,2017,48(17):3597-3604. |
| Zhang S C,Feng S N,Gu W,et al.Cloning,subcellular localization and expression analysis of a transcription factor gene SmMYB87 in Salvia miltiorrhiza[J].Chinese Traditional and Herbal Drugs,2017,48(17):3597-3604. | |
| 26 | 史文君,滕珂,陈露,等.蓝莓VcMYB启动子克隆及其功能初步分析[J].华北农学报,2017,32(2):14-20. |
| Shi W J,Teng K,Chen L,et al.Cloning and functional analysis of VcMYB promoter from Vaccinium corymbosum[J].Acta Agriculturae Boreali-Sinica,2017,32(2):14-20. | |
| 27 | 郭佳茹,胡雪虹,杨郁文,等.海岛棉MYB转录因子GbMYB5基因启动子的克隆及其在拟南芥中的表达特性分析[J].江苏农业学报,2012,28(5):957-962. |
| Guo J R,Hu X H,Yang Y W,et al.Cloning of MYB transcription factor gene(GbMYB5) promoter from Gossypium barbadense L. and functional expression characterization in transgenic Arabidopsis thaliana[J].Jiangsu Journal of Agricultural Sciences,2012,28(5):957-962. | |
| 28 | Prabu G,Theertha Prasad D.Functional characterization of sugarcane MYB transcription factor gene promoter(PScMYBAS1) in response to abiotic stresses and hormones[J].Plant Cell Reports,2012,31(4):661-669. |
| 29 | Xu F C,Liu H L,Xu Y Y,et al.Heterogeneous expression of the cotton R2R3-MYB transcription factor GbMYB60 increases salt sensitivity in transgenic Arabidopsis[J].Plant Cell,Tissue and Organ Culture (PCTOC),2018,133(1):15-25. |
| 30 | 余婧,郭玉双,雷波,等.烟草NtMYB44b转录因子基因克隆以及生物信息学和表达分析[J].植物生理学报,2020,56(2):200-208. |
| Yu J,Guo Y S,Lei B,et al.Cloning and bioinformatic and expression analyses of NtMYB44b transcription factor gene in Nicotiana tabacum[J].Plant Physiology Journal,2020,56(2):200-208. | |
| 31 | Fang Q,Wang Q,Mao H,et al.AtDIV2,an R-R-type MYB transcription factor of Arabidopsis,negatively regulates salt stress by modulating ABA signaling[J].Plant Cell Reports,2018,37(11):1499-1511. |
| 32 | Shan H,Chen S M,Jiang J F,et al.Heterologous expression of the chrysanthemum R2R3-MYB transcription factor CmMYB2 enhances drought and salinity tolerance,increases hypersensitivity to ABA and delays flowering in Arabidopsis thaliana[J].Molecular Biotechnology,2012,51(2):160-173. |
| [1] | Jiayu CAO, Lina CAO, Qiaoyi ZHANG, Shuang ZHANG, Zhibao HU, Tangrui ZHAO, Zhiru XU, Chunming LI, Xiankui QUAN, Guanjun LIU. Cloning and Expression Activity Analysis of Rubisco Small Subunit Gene Promoter in Populus × xiaohei [J]. Bulletin of Botanical Research, 2024, 44(4): 625-633. |
| [2] | ZHU De-Cai, HU Xiao-Qing, ZHAO Tong, TIAN Jing, ZHANG Yong, LIU Xue-Mei. Cloning and Expression Analysis of Promoter of CESA7 Gene in Betula platyphylla [J]. Bulletin of Botanical Research, 2018, 38(4): 551-558. |
| [3] | LI Xi-Yan, ZHAO Kai, ZHANG Xue-Mei, ZHOU Bo-Ru. Expression Analysis of Poplar MYB Transcription Factor Gene Family in Response to Salt Stress [J]. Bulletin of Botanical Research, 2017, 37(3): 424-431. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||