Bulletin of Botanical Research ›› 2021, Vol. 41 ›› Issue (4): 573-587.doi: 10.7525/j.issn.1673-5102.2021.04.013
• Research report • Previous Articles Next Articles
Bo LI, He-Xia LIU, Qin LIU, Xing-Wen ZHOU( )
)
Received:2020-06-02
Online:2021-07-20
Published:2021-03-24
Contact:
Xing-Wen ZHOU
E-mail:xingwenzhou2003@163.com
About author:LI Bo(1981—),male,PhD,associate professor,mainly engaged in plant resources utilization research.
Supported by:CLC Number:
Bo LI, He-Xia LIU, Qin LIU, Xing-Wen ZHOU. Transcriptome Analysis of Petals and to Explore Related Genes in Petal Development of Camellia nitidissima[J]. Bulletin of Botanical Research, 2021, 41(4): 573-587.
Add to citation manager EndNote|Ris|BibTeX
URL: https://bbr.nefu.edu.cn/EN/10.7525/j.issn.1673-5102.2021.04.013

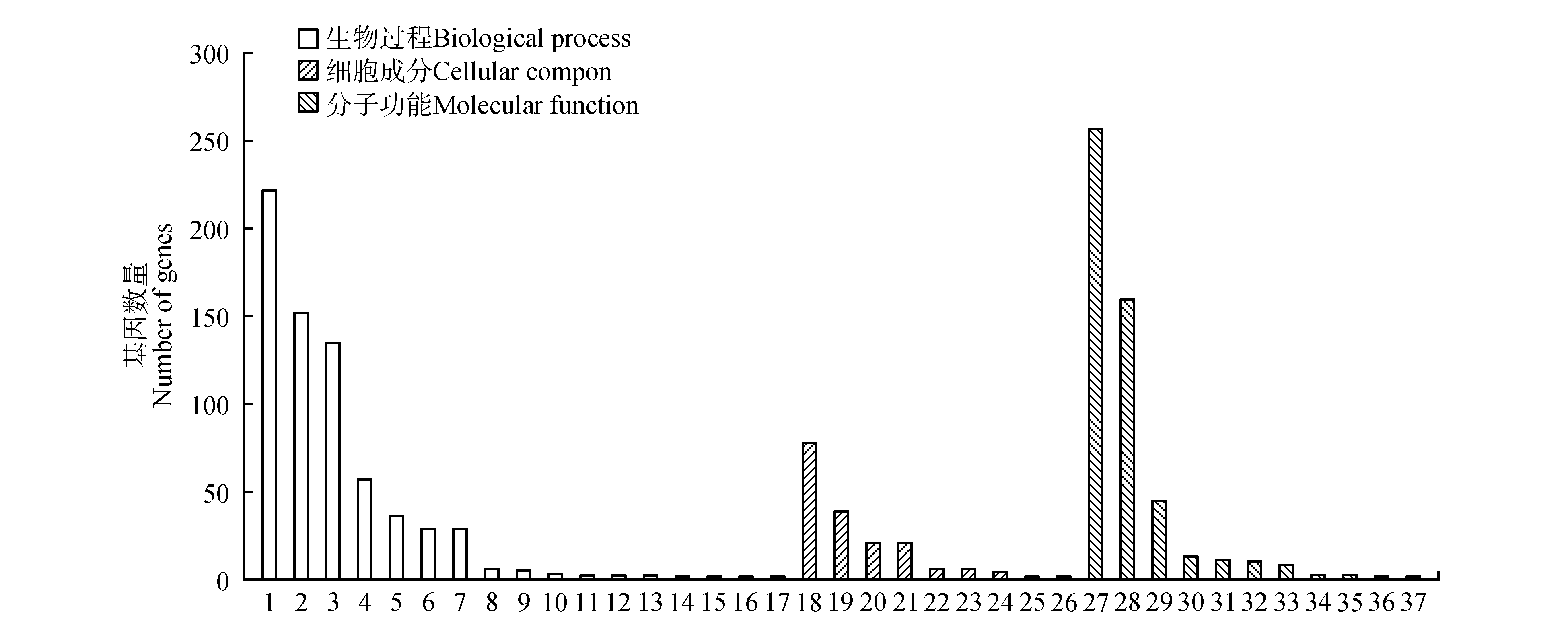
Fig.3
GO classification of DEGs in C. nitidissima1.Metabolic process;2.Single-organism process;3.Cellular process;4.Localization;5.Response to stimulus;6.Regulation of biological process;7.Biological regulation;8.Signaling;9.Cellular component organization or biogenesis;10.Multicellular organismal process;11.Growth;12.Developmental process;13.Locomotion;14.Immune system process;15.Negative regulation of biological process;16.Reproduction;17.Reproductive process;18.Catalytic activity;19:Binding;20: Transporter activity;21.Molecular function regulator;22.Nucleic acid binding transcription factor activity;23.Electron carrier activity;24.Antioxidant activity;25.Signal transducer activity;26.Structural molecule activity;27.Nutrient reservoir activity;28.Molecular transducer activity;29.Membrane;30.Membrane part;31.Cell;32.Cell part;33.Macromolecular complex;34.Organelle;35.Extracellular region;36.Supramolecular fiber;37.Organelle part
| 1 | 黄瑞斌,和太平,庄嘉,等.广西防城港市金花茶组植物资源及其保育对策[J].广西农业生物科学,2007,26(S1):32-37. |
| Huang R B,He T P,Zhuang J,et al.Plant resource and it’s conservational countermeasure of Sect.Chrysantha Chang in Fangchenggang[J].Journal of Guangxi Agricultural and Biological Science,2007,26(S1):32-37. | |
| 2 | 孔桂菊,袁胜涛,孙立.金花茶药理作用研究进展[J].时珍国医国药,2016,27(6):1459-1461. |
| Kong G J,Yuan S T,Sun L.Progress in pharmacological action research of Camellia nitidissima[J].Lishizhen Medicine and Materia Medica Research,2016,27(6):1459-1461. | |
| 3 | Dai L,Li J L,Liang X Q,et al.Flowers of Camellia nitidissima cause growth inhibition,cell-cycle dysregulation and apoptosis in a human esophageal squamous cell carcinoma cell line[J].Molecular Medicine Reports,2016,14(2):1117-1122. |
| 4 | Wang W X,Liu H Y,Wang Z N,et al.Phytochemicals from Camellia nitidissima Chi inhibited the formation of advanced glycation end-products by scavenging methylglyoxal[J].Food Chemistry,2016,205:204-211. |
| 5 | He D Y,Li X Y,Sai X,et al.Camellia nitidissima C.W.Chi:a review of botany,chemistry,and pharmacology[J].Phytochemistry Reviews,2017,17(2):327-349. |
| 6 | Fenster C B,Armbruster W S,Wilson P,et al.Pollination syndromes and floral specialization[J].Annual Review of Ecology Evolution & Systematics,2004,35:375-403. |
| 7 | Stuurman J,Hoballah M E,Broger L,et al.Dissection of floral pollination syndromes in petunia[J].Genetics,2004,168(3):1585-1599. |
| 8 | Nishihara M,Nakatsuka T.Genetic engineering of flavonoid pigments to modify flower color in floricultural plants[J].Biotechnology Letters,2011,33(3):433-441. |
| 9 | Irish V F.The Arabidopsis petal:a model for plant organogenesis[J].Trends in Plant Science,2008,13(8):430-436. |
| 10 | Abraham M C,Metheetrairut C,Irish V F.Natural variation identifies multiple loci controlling petal shape and size in Arabidopsis thaliana[J].PLoS One,2013,8(2):e56743. |
| 11 | Schmid M,Davison T S,Henz S R,et al.A gene expression map of Arabidopsis thaliana development[J].Nature Genetics,2005,37(5):501-506. |
| 12 | Han Y,Wan H H,Cheng T R,et al.Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis[J].Scientific Reports,2017,7(1):43382. |
| 13 | Wang J J,Wang H B,Ding L,et al.Transcriptomic and hormone analyses reveal mechanisms underlying petal elongation in Chrysanthemum morifolium ‘Jinba’[J].Plant Molecular Biology,2017,93(6):593-606. |
| 14 | Zhou X W,Li J Y,Zhu Y L,et al.De novo assembly of the Camellia nitidissima Transcriptome reveals key genes of flower pigment biosynthesis[J].Frontiers in Plant Science,2017,8:1545. |
| 15 | Grabherr M G,Haas B J,Yassour M,et al.Full-length transcriptome assembly from RNA-Seq data without a reference genome[J].Nature Biotechnology,2011,29(7):644-652. |
| 16 | Li B,Dewey C N.RSEM:accurate transcript quantification from RNA-Seq data with or without a reference genome[J].BMC Bioinformatics,2011,12(1):323. |
| 17 | Trapnell C,Williams B A,Pertea G,et al.Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation[J].Nature Biotechnology,2010,28(5):511-515. |
| 18 | Anders S,Huber W.Differential expression analysis for sequence count data[J].Genome Biology,2010,11(10):R106. |
| 19 | Young M D,Wakefield M J,Smyth G K,et al.Gene ontology analysis for RNA-seq:accounting for selection bias[J].Genome Biology,2010,11(2):R14. |
| 20 | Mao X Z,Cai T,Olyarchuk J G,et al.Automated genome annotation and pathway identification using the KEGG Orthology(KO) as a controlled vocabulary[J].Bioinformatics,2005,21(19):3787-3793. |
| 21 | Jason E,Bar-Joseph Z.STEM:a tool for the analysis of short time series gene expression data[J].BMC Bioinformatics,2006,7(1):191. |
| 22 | 张欢,杨英杰,李鼎立,等.杜梨根茎叶特异表达基因的RNA-Seq分析[J].园艺学报,2018,45(10):1881-1894. |
| Zhang H,Yang Y J,Li D L,et al.RNA-Seq analysis of the tissue-specific expressed genes of Pyrusbetulaefolia in root,stem and leaf[J].Acta Horticulturae Sinica,2018,45(10):1881-1894. | |
| 23 | Livak K J,Schmittgen T D.Analysis of relative gene expression data using real-time quantitative PCR and the 2-△△CT method[J].Methods,2001,25(4):402-408. |
| 24 | Powell A E,Lenhard M.Control of organ size in plants[J].Current Biology,2012,22(9):R360-R367. |
| 25 | Winship L J,Obermeyer G,Geitmann A,et al.Under pressure,cell walls set the pace[J].Trends in Plant Science,2010,15(7):363-369. |
| 26 | Buchanan B B,Gruissem W,Jones R L.Biochemistry and molecular biology of plants[M].New Jersey:Wiley,2002:85-86. |
| 27 | Cosgrove D J.Growth of the plant cell wall[J].Nature Reviews Molecular Cell Biology,2005,6(11):850-861. |
| 28 | Rose J K,Braam J,Fry S C,et al.The XTH family of enzymes involved in xyloglucan endotransglucosylation and endohydrolysis:current perspectives and a new unifying nomenclature[J].Plant & Cell Physiology,2002,43(12):1421-1435. |
| 29 | Lang X A,Li N,Li L F,et al.Integrated metabolome and transcriptome analysis uncovers the role of anthocyanin metabolism in Michelia maudiae[J].International Journal of Genomics,2019,2019:4393905. |
| 30 | Li H,Li J R,Dong Y M,et al.Time-series transcriptome provides insights into the gene regulation network involved in the volatile terpenoid metabolism during the flower development of lavender[J].BMC Plant Biology,2019,19:313. |
| 31 | Rothenberg D O,Yang H J,Chen M B,et al.Metabolome and transcriptome sequencing analysis reveals anthocyanin metabolism in pink flowers of anthocyanin-rich tea(Camellia sinensis)[J].Molecules,2019,24(6):1064. |
| 32 | Yue J Y,Zhu C X,Zhou Y,et al.Transcriptome analysis of differentially expressed unigenes involved in flavonoid biosynthesis during flower development of Chrysanthemum morifolium ‘Chuju’[J].Scientific Reports,2018,8(1):13414. |
| 33 | Singh A P,Tripathi S K,Nath P,et al.Petal abscission in rose is associated with the differential expression of two ethylene-responsive xyloglucan endotransglucosylase/hydrolase genes,RbXTH1 and RbXTH2[J].Journal of Experimental Botany,2011,62(14):5091-5103. |
| 34 | Becnel J,Natarajan M,Kipp A,et al.Developmental expression patterns of Arabidopsis XTH genes reported by transgenes and genevestigator[J].Plant Molecular Biology,2006,61(3):451-467. |
| 35 | Kurasawa K,Matsui A,Yokoyama R,et al.The AtXTH28 gene,a xyloglucan endotransglucosylase/hydrolase,is involved in automatic self-pollination in Arabidopsis thaliana[J].Plant & Cell Physiology,2009,50(2):413-422. |
| 36 | Ariizumi T,Amagai M,Shibata D,et al.Comparative study of promoter activity of three anther-specific genes encoding lipid transfer protein,xyloglucan endotransglucosylase/hydrolase and polygalacturonase in transgenic Arabidopsis thaliana[J].Plant Cell Reports,2002,21(1):90-96. |
| 37 | Harada T,Torii Y,Morita S,et al.Cloning,characterization,and expression of xyloglucan endotransglucosylase/hydrolase and expansin genes associated with petal growth and development during carnation flower opening[J].Journal of Experimental Botany,2011,62(2):815-823. |
| 38 | Santner A,Estelle M.Recent advances and emerging trends in plant hormone signalling[J].Nature,2009,459(7250):1071-1078. |
| 39 | Han Y,Yong X,Yu J Y,et al.Identification of candidate adaxial-abaxial-related genes regulating petal expansion during flower opening in Rosa chinensis “Old Blush”[J].Frontiers in Plant Science,2019,10:1098. |
| 40 | Swarup K,Benková E,Swarup R,et al.The auxin influx carrier LAX3 promotes lateral root emergence[J].Nature Cell Biology,2008,10(8):946-954. |
| 41 | Stepanova A N,Robertson-Hoyt J,Yun J,et al.TAA1-Mediated auxin biosynthesis is essential for hormone crosstalk and plant development[J].Cell,2008,133(1):177-191. |
| 42 | Riechmann J L,Ratcliffe O J.A genomic perspective on plant transcription factors[J].Current Opinion in Plant Biology,2000,3(5):423-434. |
| 43 | Zhang Y F,Cao G Y,Qu L J,et al.Characterization of Arabidopsis MYB transcription factor gene AtMYB17 and its possible regulation by LEAFY and AGL15[J].Journal of Genetics and Genomics,2009,36(2):99-107. |
| 44 | Pastore J J,Limpuangthip A,Yamaguchi N,et al.Late meristem IDENTITY2 acts together with leafy to activate APETALA1[J].Development,2011,138(15):3189-3198. |
| 45 | Lee D K,Geisler M,Springer P S.LATERAL ORGAN FUSION1 and LATERAL ORGAN FUSION2 function in lateral organ separation and axillary meristem formation in Arabidopsis[J].Development,2009,136(14):2423-2432. |
| 46 | Szécsi J,Joly C,Bordji K,et al.BIGPETALp,a bHLH transcription factor is involved in the control of Arabidopsis petal size[J].The EMBO Journal,2006,25(16):3912-3920. |
| 47 | 张璐,杜相革.植物水孔蛋白研究进展[J].植物科学学报,2014,32(3):304-314. |
| Zhang L,Du X G.Recent advances in plant aquaporins[J].Plant Science Journal,2014,32(3):304-314. | |
| 48 | Ma N,Xue J Q,Li Y H,et al.Rh-PIP2;1,a Rose aquaporin gene,is involved in ethylene-regulated petal expansion[J].Plant Physiology,2008,148(2):894-907. |
| 49 | 雍伟东,谭克辉,许智宏,等.高等植物开花时间决定的基因调控研究[J].科学通报,2000,45(5):455-466. |
| Yong W D,Chong K,Xu Z H,et al.Gene control of flowering time in higher plants[J].Chinese Science Bulletin,2000,45(18):1633-1642. | |
| 50 | Blümel M,Dally N,Jung C.Flowering time regulation in crops-what did we learn from Arabidopsis?[J].Current Opinion in Biotechnology,2015,32:121-129. |
| 51 | Bouchée F,Lobet G,Tocquin P,et al.FLOR-ID:an interactive database of flowering-time gene networks in Arabidopsis thaliana[J].Nucleic Acids Research,2016,44(D1):D1167-D1171. |
| 52 | Wang S,Huang H J,Han R,et al.BpAP1 directly regulates BpDEF to promote male inflorescence formation in Betula platyphylla × B.pendula[J].Tree Physiology,2019,39(6):1046-1060. |
| 53 | Agrawal G K,Abe K,Yamazaki M,et al.Conservation of the E function for floral organ identity in rice revealed by the analysis of tissue culture induced loss of function mutants of the OsMADS1 gene[J].Plant Molecular Biology,2005,59(1):125-135. |
| 54 | Theissen G.Development of floral organ identity:stories from the MADS house[J].Current Opinion in Plant Biology,2001,4(1):75-85. |
| 55 | Favaro R,Pinyopich A,Battaglia R,et al.MADS-box protein complexes control carpel and ovule development in Arabidopsis[J].The Plant Cell,2003,15(11):2603-2611. |
| 56 | Sablowski R W M,Meyerowitz E M.A homolog of NO APICAL MERISTEM is an immediate target of the floral homeotic genes APETALA3/PISTILLATA[J].Cell,1998,92(1):93-103. |
| 57 | Ma H,Depamphilis C.The ABCs of floral evolution[J].Cell,2000,101(1):5-8. |
| 58 | Fei Y,Liu Z X.Isolation and characterization of the PISTILLATA Ortholog gene from Cymbidium faberi Rolfe[J].Agronomy,2019,9(8):425. |
| [1] | ZHU Li-Li, DU Qing-Xin, HE Feng, QING Jun, DU Hong-Yan. Sequencing Analysis of Transcriptome of Male Floral Bud at Two Development Stages in Eucommia ulmoides [J]. Bulletin of Botanical Research, 2020, 40(2): 284-292. |
| [2] | ZHANG Li-Li, ZHANG Fu-Chun. Transcriptomic Analysis of the Halostachys caspica in Response to Short-term Salt Stress [J]. Bulletin of Botanical Research, 2018, 38(1): 91-99. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||