Bulletin of Botanical Research ›› 2026, Vol. 46 ›› Issue (2): 195-208.doi: 10.7525/j.issn.1673-5102.2026.02.001
• Original Paper • Next Articles
Wenbin DING, Yuehan HU, Yinjia BAI, Yuhan YANG, Hua XUE( )
)
Received:2025-12-24
Online:2026-03-20
Published:2026-04-02
Contact:
Hua XUE
E-mail:xuehua2013@bjfu.edu.cn
CLC Number:
Wenbin DING, Yuehan HU, Yinjia BAI, Yuhan YANG, Hua XUE. Expression Characteristics and Functional Analysis of PtNF-Ys Gene during Early Flower Development in Pinus tabuliformis[J]. Bulletin of Botanical Research, 2026, 46(2): 195-208.
Add to citation manager EndNote|Ris|BibTeX
URL: https://bbr.nefu.edu.cn/EN/10.7525/j.issn.1673-5102.2026.02.001
Table 1
The primers information
基因名称 Gene name | 正向引物序列(5′→3′) Forward primer sequence (5′→3′) | 反向引物序列(5′→3′) Reverse primer sequence(5′→3′) |
|---|---|---|
| RqPtNF-YB8 | ATGGCAGACTTAGCGAGCTCCG | GAGGACTTTCTTGGCTGGTGA |
| RqPtNF-YC1 | ATGGACCACCACAACCACC | CCTGCTGCCCTCCACTCT |
| RqPtTFL2 | ATGGCCAGGTTCAGAGAGCCT | ACATCGCCAATGACTCTTCCT |
| RqPtCOL1 | ATGGTGAAGGAGGAGGATTGT | CCTTCACGATTCCTGCCTC |
| PtNF-YB8 | ATGGCAGACTTAGCGAGCTCC | CCTAATATTTGGTAATGCTCCAAAA |
| PtNF-YB8-AD | CCATGGAGGCCAGTGAATTCATGGCAGACTTAGC GAGCTCC | CCATGGAGGCCAGTGAATTCCCTAATATTTGGTAATG CTCCAAAA |
| PtNF-YC4-BD | TGGCCATGGAGGCCGAATTCATGCATGATTTAGAA CAGACATCAGAC | CGCTGCAGGTCGACGGATCCCTAACTGCTAGAGGAT GGGCC |
| PtNF-YC10-BD | TGGCCATGGAGGCCGAATTCATGCATGATATAGAA CAGAC | CGCTGCAGGTCGACGGATCCCTATGCTGCTGGATGC |
| PtNF-YC9-BD | TGGCCATGGAGGCCGAATTCATGAGGAAGAAGCT CGATACCC | CGCTGCAGGTCGACGGATCCTATCCATCATCATTGTC ATAGTCATCCTC |
| PtNF-YA3-BD | TGGCCATGGAGGCCGAATTCATGACCGTTCGGGA ATTTTTGC | CGCTGCAGGTCGACGGATCCTCACTGGATCACAACA ACTCGT |
| PtNF-YA9-BD | TGGCCATGGAGGCCGAATTCATGGAGAACACGCC TACACATAC | CGCTGCAGGTCGACGGATCCCTAGGAATCCGTGTTC TGCTGT |
| PtNF-YB8-1300 | AACACGGGGGACGAGCTCGGTACCATGGCAGAC TTAGCGAGCTCC | TTCTTCTCCTTTGCCCATGTCGACTCTAGACCTAATAT TTGGTAATGCTCCAAAA |
| PtActin | GGCTGACACCATCACCAGAATC | GTTGGTCGCCCTCGTCATACT |
| AtActin | CGTATGAGCAAAGAGATCAC | CACATCTGTTGGAAGGTGCT |
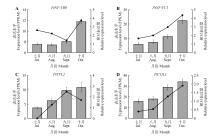

Fig.3
Expression patterns and relative transcript levels of PtTFL2,PtCOL1,PtNF-YB8,and PtNF-YC1 genesduring the early stages of flower bud differentiationThe line charts represented transcript abundance(FPKM) obtained from transcriptome sequencing, while bar charts indicated relative expression levels determined by qRT-PCR. All samples were collected from shoot apical meristems at corresponding developmental stages. Each data point represented the mean of three replicates, with error bars indicating standard deviation(SD).
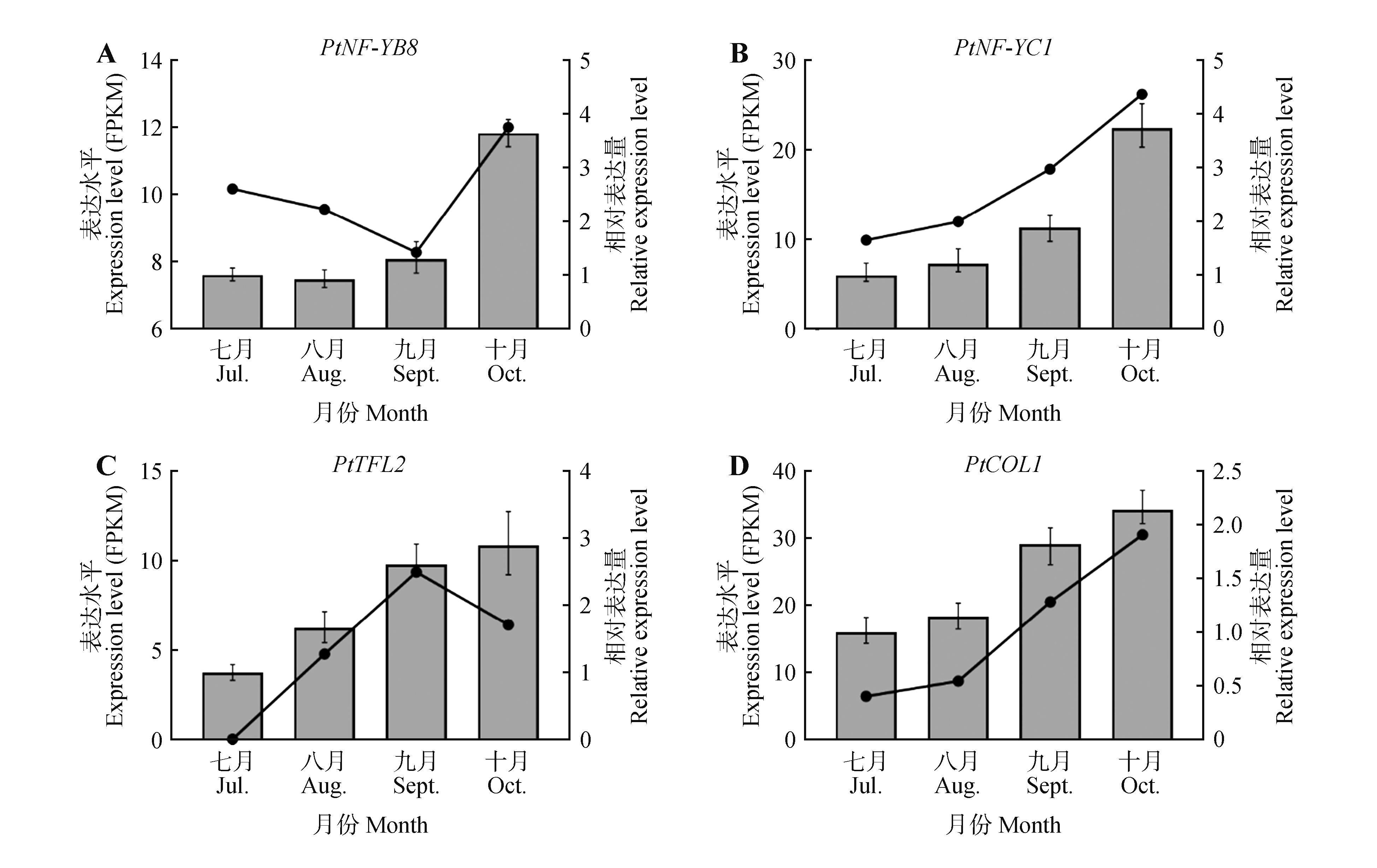
Table 2
Information of predicted PtNF - YB8 promoter elements
序号 Number | 序列 Sequence | 数量 Quantity | 位置 Location | 顺式作用元件 Cis-acting regulatory element |
|---|---|---|---|---|
| 1 | TTACTTAA | 1 | 342 | Part of a light responsive element |
| 2 | GCCACT | 2 | 420;529 | Related to meristem expression |
| 3 | CCATCTTTTT | 2 | 1 458;1 550 | Involved in salicylic acid responsiveness |
| 4 | CACGTC | 1 | 1 314 | Involved in light responsiveness |
| 5 | CAAAGATATC | 1 | 1 265 | Involved in circadian control |
| 6 | TGAGTCA | 2 | 752;943 | Involved in endosperm expression |
| 7 | ACGTG | 2 | 1 315;1 996 | Involved in the abscisic acid responsiveness |
| 8 | ATGATAAGGTC | 2 | 648;839 | Part of a light responsive element |
| 9 | AAGATAAGGCT | 1 | 1 217 | Part of a light responsive element |
| 10 | TTCCAACAAACCCC | 1 | 1 201 | Gibberellin-responsive element and part of a light responsive element |
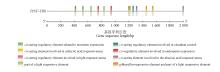

Fig.4
Schematic diagram of the predicted locations of cis-acting elements in the PtNF-YB8 promoterThe colored boxes represented different types of cis-acting regulatory elements. The promoter of PtNF-YB8 gene was depicted by horizontal lines incorporating various colored segments, with each segment signifying a different class of cis-acting elements. A numerical horizontal axis beneath the promoter indicated the distance of these elements relative to the transcription start site (unit: base pairs), while 5′ and 3′ denoted the orientation of the nucleic acid strand.
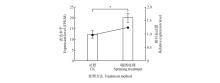
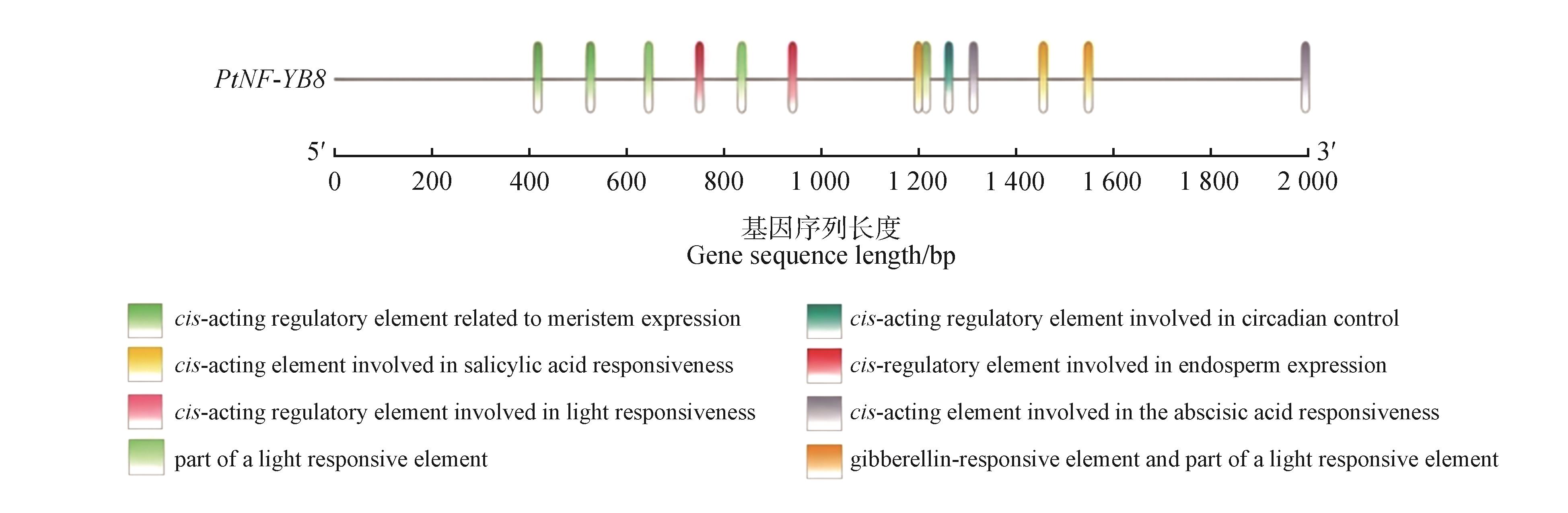

Fig.5
Expression level of PtNF-YB8 gene in shoot apical meristems of Pinus tabuliformis after gibberellin treatmentThe line charts represented transcript abundance (FPKM) obtained from transcriptome sequencing, while bar charts indicated relative expression levels determined by qRT-PCR. The ‘CK’ sample consisted of untreated shoot apical meristems collected in mid-October, while the ‘spraying treatment’ sample consisted of shoot apical meristems collected in mid-October, three months after being sprayed with gibberellin4+7. Each data point represented the mean of three replicates, with error bars indicating standard deviation (SD) . *. Significant difference at P<0.05.
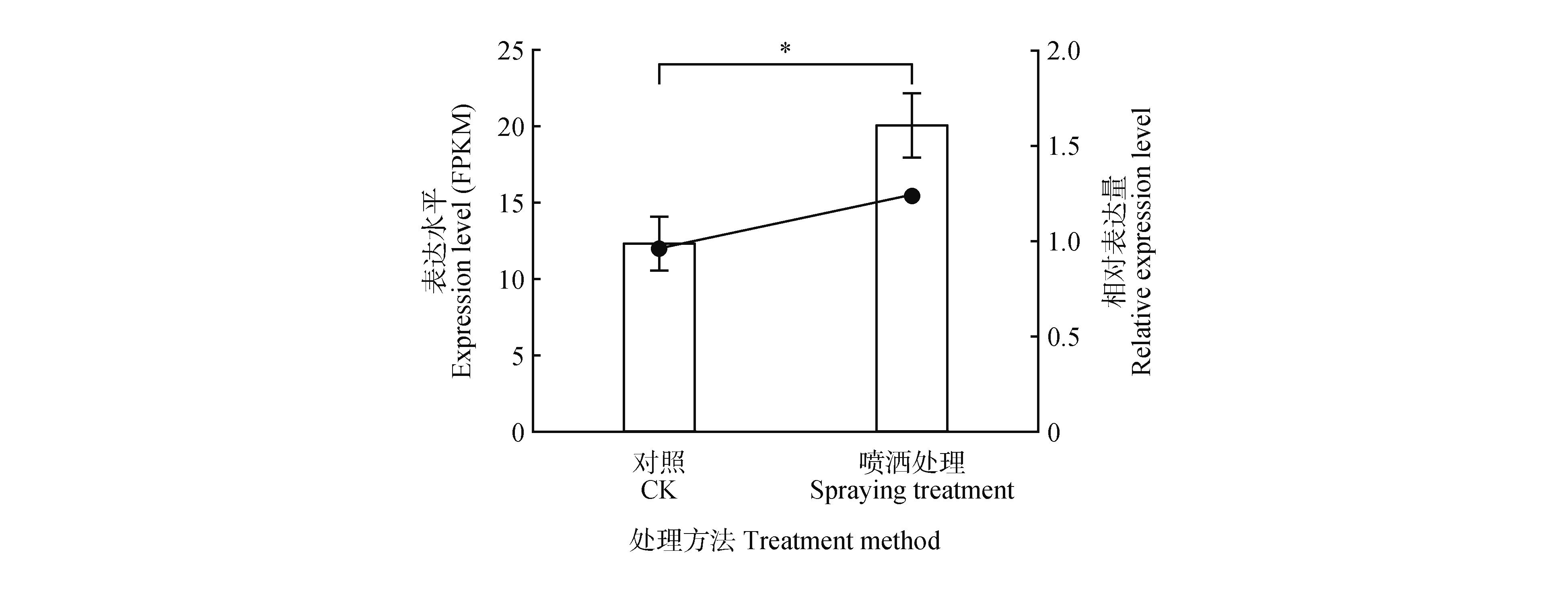
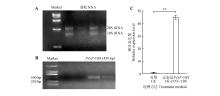
Fig.7
Clone and expression analysis of PtNF-YB8 gene in Arabidopsis transgenic linesA. Extraction and integrity analysis of total RNA from Pinus tabuliformis. Distinct 28S and 18S rRNA bands were observed via agarose gel electrophoresis, indicating that the extracted RNA possessed good integrity; B. PCR detection of transgenic Arabidopsis plants. A specific amplicon of approximately 459 bp confirmed the successful integration of PtNF-YB8 gene into the Arabidopsis genome. C. qRT-PCR analysis of PtNF-YB8 expression in confirmed transgenic lines. The expression level of PtNF-YB8 gene was significantly elevated in overexpression plants(OE-PtNF-YB8) compared with the wild type(WT), with a highly significant difference(P<0.01). Error bars represented the standard deviation(SD) of three replicates.
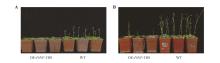
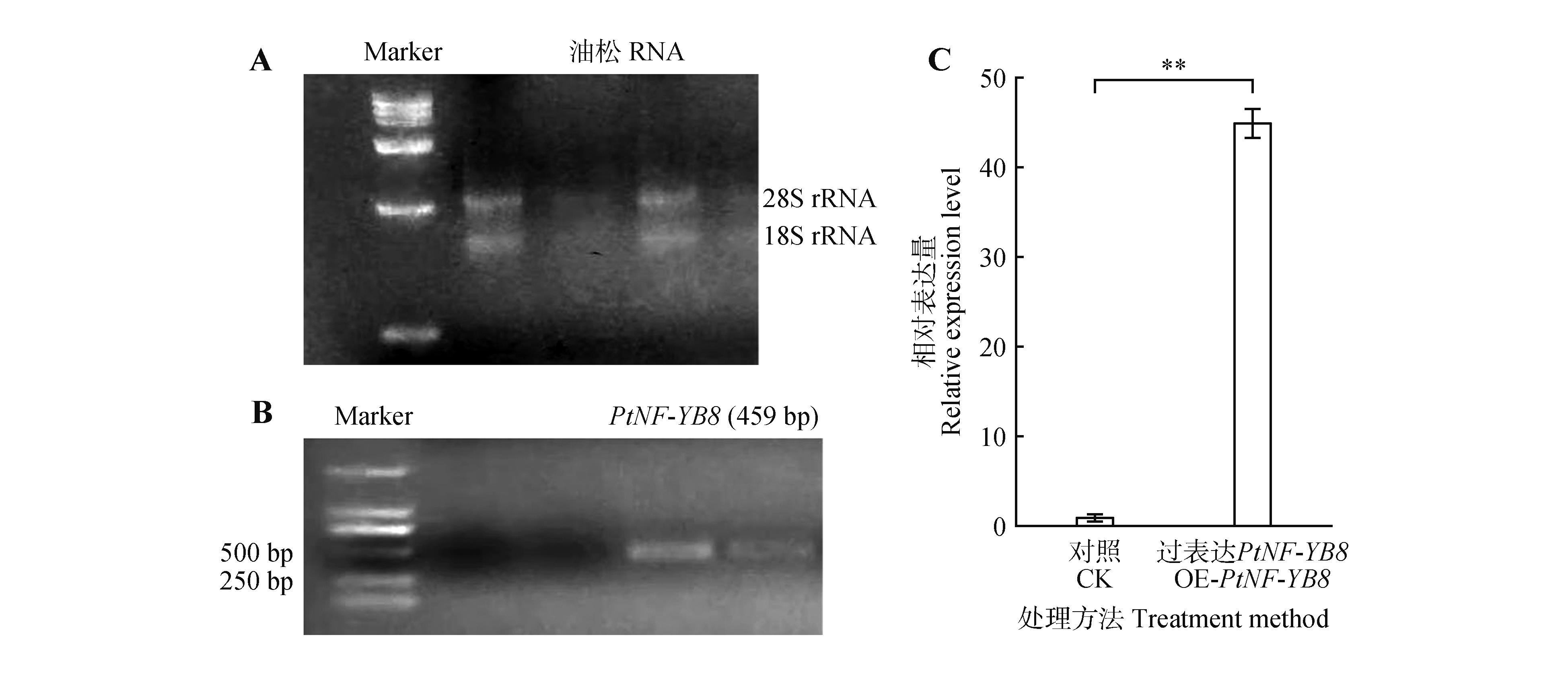

Fig.8
Late flowering phenotype of OE-PtNF-YB8 transgenic ArabidopsisPtNF-YB8 overexpression lines(OE-PtNF-YB8,left three pots) and wild-type plants(WT, right three pots) were grown under identical conditions. Photographs show representative plants at 40 d (A) or 60 d (B) since seed imbibition.
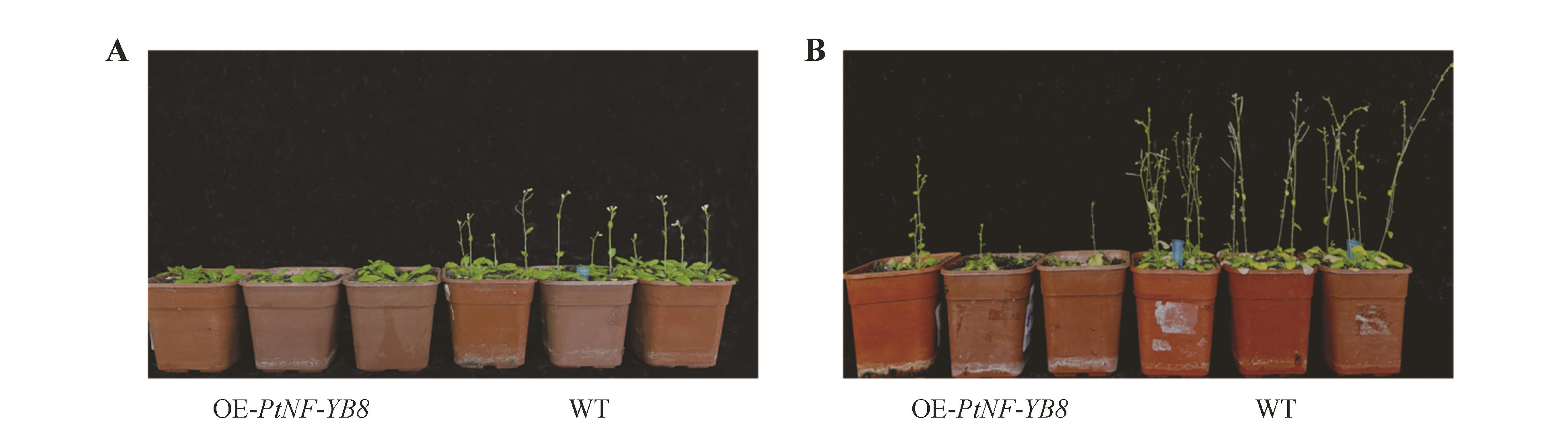
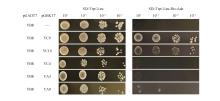


Fig.9
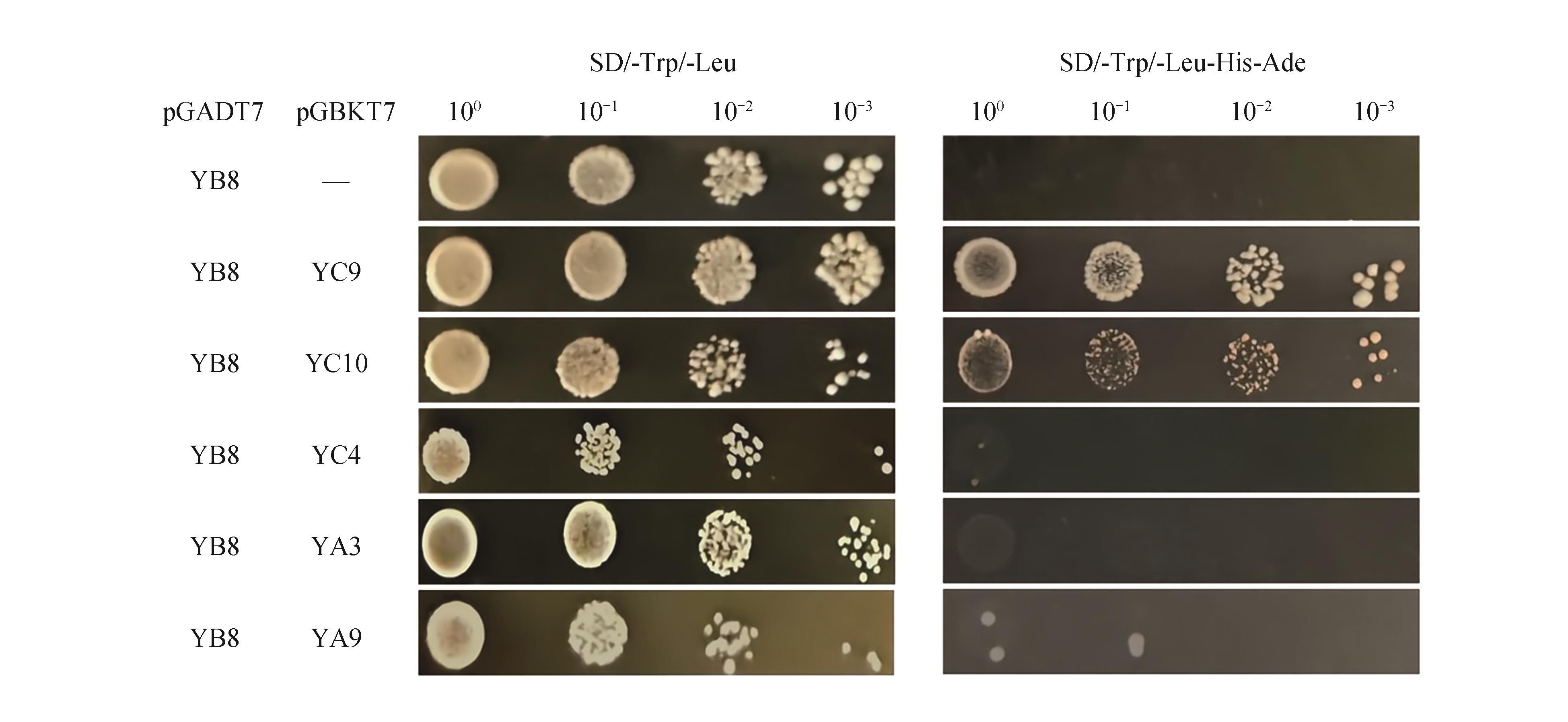
Yeast two-hybrid analysis of the interaction between PtNF-YB8 and different PtNF-Y family membersThe AH109 yeast strain was co-transformed with pGADT7-YB8 and various pGBKT7-Y constructs (YC9,YC10,YC4,YA3,YA9), followed by growth screening on selective media. SD/-Trp/-Leu medium was used to confirm successful co-transformation. SD/-Trp/-Leu/-His/-Ade medium was used to assess protein-protein interactions. Yeast suspensions were spotted at serial dilution factors (100,101,102,103).
| [1] | 樊军锋,杨培华,郭树杰,等.陕西油松遗传改良研究进展[J].西北农林科技大学学报(自然科学版),2006,34(1):45-50. |
| FAN J F, YANG P H, GUO S J,et al.Research progress of genetics and improvement on Pinus tabulaeformis Carriere of Shaanxi Province[J].Journal of Northwest A&F University(Natural Science Edition),2006,34(1):45-50. | |
| [2] | NIU S H, LI J, BO W H,et al.The Chinese pine genome and methylome unveil key features of conifer evolution[J].Cell,2022,185(1):204-217.e14. |
| [3] | 窦春蕊,李玉英,张冬红.油松种子园建立与经营技术研究现状[J].西北农业学报,2004,13(3):162-168. |
| DOU C R, LI Y Y, ZHANG D H.Summary on construction and management techniques of seed orchard of Pinus tabulaeformis [J].Acta Agriculturae Boreali-occidentalis Sinica,2004,13(3):162-168. | |
| [4] | LI W, WANG X R, LI Y,et al.Stability in and correlation between factors influencing genetic quality of seed lots in seed orchard of Pinus tabuliformis Carr.over a 12-year span[J].PLoS One,2011,6(8):e23544. |
| [5] | JARILLO J A, PIÑEIRO M.Timing is everything in plant development.The central role of floral repressors[J].Plant Science,2011,181(4):364-378. |
| [6] | MICHAELS S D, AMASINO R M.Loss of FLOWERING LOCUS C activity eliminates the late-flowering phenotype of FRIGIDA and autonomous pathway mutations but not responsiveness to vernalization[J].The Plant Cell,2001,13(4):935-941. |
| [7] | TOKUTSU R, FUJIMURA-KAMADA K, MATSUO T,et al.The CONSTANS flowering complex controls the protective response of photosynthesis in the green alga Chlamydomonas [J].Nature Communications,2019,10(1):4099. |
| [8] | HAYAMA R, SARID-KREBS L, RICHTER R,et al.PSEUDO RESPONSE REGULATORs stabilize CONSTANS protein to promote flowering in response to day length[J].The EMBO Journal,2017,36(7):904-918. |
| [9] | LÜ X C, ZENG X L, HU H M,et al.Structural insights into the multivalent binding of the Arabidopsis FLOWERING LOCUS T promoter by the CO-NF-Y master transcription factor complex[J].The Plant Cell,2021,33(4):1182-1195. |
| [10] | CHEN F, ZHANG X T, LIU X,et al.Evolutionary analysis of MIKCc-type MADS-box genes in gymnosperms and angiosperms[J].Frontiers in Plant Science,2017,8:895. |
| [11] | 刘洋,李帮同,杜桂华,等.油松FT/TFL1-like基因表达模式与调控分析[J].北京林业大学学报,2018,40(10):60-66. |
| LIU Y, LI B T, DU G H,et al.Expression profiles and regulation of FT/TFL1-like genes in Pinus tabuliformis [J].Journal of Beijing Forestry University,2018,40(10):60-66. | |
| [12] | HELLIWELL C A, ROBERTSON M, FINNEGAN E J,et al.Vernalization-repression of Arabidopsis FLC requires promoter sequences but not antisense transcripts[J].PLoS One,2011,6(6):e21513. |
| [13] | ASANTE D K A, YAKOVLEV I A, FOSSDAL C G,et al.Effect of bud burst forcing on transcript expression of selected genes in needles of Norway spruce during autumn[J].Plant Physiology and Biochemistry,2009,47(8):681-689. |
| [14] | DENNIS L, PEACOCK J.Genes directing flower development in Arabidopsis [J].The Plant Cell,2019,31(6):1192-1193. |
| [15] | SONG Y T, LIU S W, MA J J,et al.The MADS-box transcription factor DAL1 links age to reproductive development through regulation of LEAFY homologs in conifer[J].Plant Physiology,2025,198(4):kiaf139. |
| [16] | NIU S H, LI Z X, YUAN H W,et al.Transcriptome characterisation of Pinus tabuliformis and evolution of genes in the Pinus phylogeny[J].BMC Genomics,2013,14:263. |
| [17] | 石鹏,王永,张大鹏.赤霉素调控林木生长发育的研究进展[J].江西农业学报,2021,33(2):33-41. |
| SHI P, WANG Y, ZHANG D P.Advances in gibberellins regulating growth and development of tree[J].Acta Agriculturae Jiangxi,2021,33(2):33-41. | |
| [18] | ZHANG S W, ZHANG D, FAN S,et al.Effect of exogenous GA3 and its inhibitor paclobutrazol on floral formation,endogenous hormones,and flowering-associated genes in ‘Fuji’ apple(Malus domestica Borkh.)[J].Plant Physiology and Biochemistry,2016,107:178-186. |
| [19] | KONG L S, VON ADERKAS P, ZAHARIA I,et al.Analysis of phytohormone profiles during male and female cone initiation and early differentiation in long-shoot buds of lodgepole pine[J].Journal of Plant Growth Regulation,2012,31(4):478-489. |
| [20] | ROSENBERG O, ALMQVIST C, WESLIEN J,et al.Systemic insecticide and gibberellin reduced cone damage and increased flowering in a spruce seed orchard[J].Journal of Economic Entomology,2012,105(3):916-922. |
| [21] | MANTOVANI R.The molecular biology of the CCAAT-binding factor NF-Y[J].Gene,1999,239(1):15-27. |
| [22] | LALOUM T, DE MITA S, GAMAS P,et al.CCAAT-box binding transcription factors in plants:Y so many?[J].Trends in Plant Science,2013,18(3):157-166. |
| [23] | ROMIER C, COCCHIARELLA F, MANTOVANI R,et al.The NF-YB/NF-YC structure gives insight into DNA binding and transcription regulation by CCAAT factor NF-Y[J].Journal of Biological Chemistry,2003,278(2):1336-1345. |
| [24] | YU T F, LIU Y, FU J D,et al.The NF-Y-PYR module integrates the abscisic acid signal pathway to regulate plant stress tolerance[J].Plant Biotechnology Journal,2021,19(12):2589-2605. |
| [25] | GNESUTTA N, MANTOVANI R, FORNARA F.Plant flowering:imposing DNA specificity on histone-fold subunits[J].Trends in Plant Science,2018,23(4):293-301. |
| [26] | CAO S H, KUMIMOTO R W, GNESUTTA N,et al.A distal CCAAT/NUCLEAR FACTOR Y complex promotes chromatin looping at the FLOWERING LOCUS T promoter and regulates the timing of flowering in Arabidopsis [J].The Plant Cell,2014,26(3):1009-1017. |
| [27] | KIM S K, PARK H Y, JANG Y H,et al.OsNF-YC2 and OsNF-YC4 proteins inhibit flowering under long-day conditions in rice[J].Planta,2016,243(3):563-576. |
| [28] | GUO Y T, NIU S H, EL-KASSABY Y A,et al.Transcriptome-wide isolation and expression of NF-Y gene family in male cone development and hormonal treatment of Pinus tabuliformis [J].Physiologia Plantarum,2021,171(1):34-47. |
| [29] | 刘红梅,郑永涛,郭盈添,等.油松PtNF-YC1基因鉴定及其调控球花发育的作用机制研究[J].北京林业大学学报,2023,45(9):1-8. |
| LIU H M, ZHENG Y T, GUO Y T,et al.Identification of PtNF-YC1 of Pinus tabuliformis and its molecular mechanism involved in regulation of cone development[J].Journal of Beijing Forestry University,2023,45(9):1-8. | |
| [30] | WEI Q, MA C, XU Y J,et al.Control of chrysanthemum flowering through integration with an aging pathway[J].Nature Communications,2017,8(1):829. |
| [31] | YANG W B, ZHOU C C, GUO Y T,et al.Genome-wide identification of the Pinus tabuliformis CONSTANS-like gene family and their potential roles in reproductive cone development[J].International Journal of Biological Macromolecules,2024,254:127621. |
| [32] | OBESO J R.The costs of reproduction in plants[J].New Phytologist,2002,155(3):321-348. |
| [33] | ZHANG B Q, FENG M H, ZHANG J,et al.Involvement of CONSTANS-like proteins in plant flowering and abiotic stress response[J].International Journal of Molecular Sciences,2023,24(23):16585. |
| [1] | Bi QIN, Mingyang LIU, Meng WANG, Lifeng WANG, Fei HUANG. Cloning and Expression Analysis of DELLA gene HbRGL1 from Hevea brasiliensis Müll. Arg [J]. Bulletin of Botanical Research, 2022, 42(6): 997-1004. |
| [2] | Hua-Feng CHEN, Song-Le FAN, Li-Feng WANG. Structural and Functional Analysis of Gibberellin Signaling Transduction Pathway Gene HbRGA1 in Rubber Tree [J]. Bulletin of Botanical Research, 2021, 41(4): 531-539. |
| [3] | LIU Wei;XIE Bing;NI Guo-Ping;DENG Guang-Hua*. Influence of Gibberellin and Amino Acid on Branch and Leaf Growth of Syzygium grijsii [J]. Bulletin of Botanical Research, 2011, 31(2): 218-226. |
| [4] | HU Shang-Lian;JIA Ju-Qing;CAO Ying;SUN Xia;LU Xue-Qin. Effect of GA3 and IAA on the Dynamic Accumulation of Holocellulose and Chlorophyl Content in Neosinocalamus affinis [J]. Bulletin of Botanical Research, 2008, 28(6): 737-740. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||